ISSN: 1839-9940
J Genomics 2016; 4:7-9. doi:10.7150/jgen.14456 This volume Cite
Short Research Communication
Draft Genome Sequences of Two Pathogenic Corynebacterial Species Isolated from Cows
1. Institute of Biological Sciences, Federal University of Pará (UFPA), Belém, PA, Brazil.
2. Institute of Biological Sciences, Federal University of Minas Gerais (UFMG), Belo Horizonte, MG, Brazil.
3. National Reference Laboratory for Aquatic Animal Diseases, Ministry of Fisheries and Aquaculture, Federal University of Minas Gerais (UFMG), Belo Horizonte, MG, Brazil.
4. Unité de Prévention et Thérapie Moléculaires des Maladies Humaines, Institut Pasteur, Paris, France.
Published 2016-3-2
Abstract
The species Corynebacterium renale, Corynebacterium pilosum, and Corynebacterium cystitidis were initially thought to be the same species C. renale, but with different immunological types. These bacteria are the causative agent of cystitis, urethritis and pyelonephritis and are found usually as constituents of the normal flora in the lower urogenital tract of cattle. Therefore, we present the draft genome sequences of two pathogenic Corynebacterium species: C. renale CIP 52.96 and C. pilosum CIP 103422. The genome sequences of these species have 2,322,762 bp with 2,218 protein encoding genes and 2,548,014 bp with 2,428 protein encoding genes, respectively. These genomes can help clarify the virulence mechanisms of these unknown bacteria and enable the development of more effective methods for control.
Keywords: Corynebacterium spp., Genome Sequencing, Draft Genomes, Ion Torrent.
Introduction
The Corynebacterium genus was created to include the pathogenic species Corynebacterium diphtheriae, the causative agent of diphtheria (1). Currently, approximately 90 species with various lifestyles comprise this genus, including human, animal and plant pathogens (2).
The species Corynebacterium renale, a bacterium of the Corynebacterium genus, has not been well studied. These bacteria were first described with three immunological types. Only after the development of genetic analyses and chemotaxonomic methods was it possible to distinguish this group from the following three species: Corynebacterium renale (Type I), Corynebacterium pilosum (Type II), and Corynebacterium cystitidis (Type III) (3).
These corynebacteria groups are usually found as constituents of the normal flora in the lower urogenital tract, and their presence is thought to be a precondition for the development of cystitis, urethritis and pyelonephritis in cattle (4,5). Their adhesion to urinary epithelial cells is mediated by pili structures present on the bacteria cell surface and is recognized as an important virulence factor.
In this report, we announce the draft genome sequences of the following two corynebacteria pathogenic species: (i) C. pilosum strain CIP 103422 isolated from cow urine in Japan and (ii) C. renale strain CIP 52.96 isolated from a cow in the United Kingdom. At the time this work was prepared, no other genome assembly was publicly available for the species C. renale.
The genome sequencing of the isolates was performed by the Ion Torrent PGM (Personal Genome Machine) platform (Life Technologies), using a fragment library. The de novo assembly of the sequences into contigs was achieved using MIRA (6), and gap closure was performed with the Lasergene v.11 Suite (DNASTAR). The assembly produced a total of 19 contigs for C. pilosum CIP103422 with 2,548,014 total base pairs and 11 contigs for C. renale CIP 52.96 with 2,322,762 total base pairs and a G+C content of 60.7% and 59.1%, respectively. Automatic annotation using the RAST server (7) allowed for the identification of 2,387 Coding DNA Sequences (CDSs) in the C. pilosum CIP103422 and 2,218 CDSs in the C. renale CIP 52.96; 61 RNA genes were predicted for both genomes.
Additionally, we used the SEED annotation environment (http://www.theseed.org/) to investigate the genomic basis of several biochemical features that differed between corynebacterial isolates. The presence of genes coding for enzymes involved in invasion and intracellular resistance, such as L-aspartate oxidase (EC 1.4.3.16) and Quinolinate synthetase (EC 2.5.1.72), were only identified in C. renale CIP 52.96. A distinctive characteristic of C. pilosum CIP103422 was the presence of potential activity for respiratory nitrate reductase (EC 1.7.99.4), which was not detected in C. renale CIP 52.96.
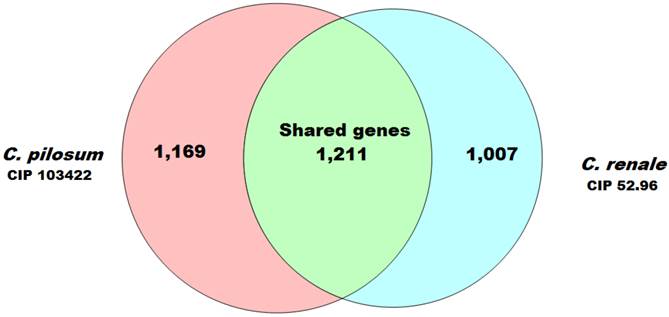
Furthermore, analyses using PGAP software (8) with the MultiParanoid (MP) method with ≥50% of coverage and ≥50% of identity allowed the prediction of 1211 CDSs that were shared between both genomes. In these genes, CDS usually plays a crucial role in the maintenance of key aspects for the organism's biology and is a great target for drug development (9). Additionally, we predicted 1169 CDSs exclusive for C. pilosum CIP103422 and 1007 CDSs for C. renale CIP 52.96. These genes could confer an adaptive advantage and niche adaptation (10) (Fig. 1).
Venn diagram demonstrating the numbers of coding sequences shared by the two Corynebacterium species analyzed in this study.
The genome projects have been deposited in GenBank under the following accession numbers: NZ_LDYE00000000.1 (C. renale CIP 52.96) and NZ_LDYD00000000.1 (C. pilosum CIP103422.
Acknowledgements
This work was supported by the Coordination for the Improvement of Higher Education Personnel (CAPES) and the Ministry of Fishing and Aquaculture. We also acknowledge support from the National Council for Scientific and Technological Development (CNPq) and the Genome and Proteome Network of the State of Pará (RPGP).
Competing Interests
The authors have declared that no competing interest exists.
References
1. Cerdeño-Tárraga AM, Efstratiou A, Dover LG, Holden MTG, Pallen M, Bentley SD. et al. The complete genome sequence and analysis of Corynebacterium diphtheriae NCTC13129. Nucleic Acids Res. 2003Nov15;31(22 ):6516-23
2. Tauch A, Sandbote J. The Family Corynebacteriaceae. In: (ed.) Rosenberg E, DeLong E, Lory S, Stackebrandt E, Thompson F. The Prokaryotes SE - 187. Springer Berlin Heidelberg. 2014:239-77
3. Winn WC, Allen SD, Janda WM, Koneman EW, Schreckenberger PC, Procop GW. et al. Koneman's Color Atlas and Textbook of Diagnostic Microbiology; 6th ed; Koneman EW, editor. Philadelphia: Lippincott Williams & Wilkins. 2005
4. Sato H, Yanagawa R, Fukuyama H. Adhesion of Corynebacterium renale, Corynebacterium pilosum, and Corynebacterium cystitidis to bovine urinary bladder epithelial cells of various ages and levels of differentiation. Infect Immun. 1982;36(3):1242-5
5. Yeruham I, Elad D, Perl S, Avidar Y, Israeli B, Shlosberg A. Isolation of Corynebacterium pilosum and Actinomyces pyogenes from cystitis and vulvovaginitis infection in a 2-month-old female calf. Zentralbl Veterinarmed B. 1999;46(2):127-30
6. Chevreux B, Wetter T, Suhai S. Genome Sequence Assembly Using Trace Signals and Additional Sequence Information. Comput Sci Biol Proc Ger Conf Bioinforma. 1999(1995):45-56
7. Aziz RK, Bartels D, Best A, DeJongh M, Disz T, Edwards R. et al. The RAST Server: rapid annotations using subsystems technology. BMC Genomics. 2008;9:75
8. Zhao Y, Wu J, Yang J, Sun S, Xiao J, Yu J. PGAP: Pan-genomes analysis pipeline. Bioinformatics. 2012;28(3):416-8
9. Medini D, Donati C, Tettelin H, Masignani V, Rappuoli R. The microbial pan-genome. Curr Opin Genet Dev. 2005;15(6):589-94
10. Rasko D, Rosovitz MJ, Myers GS, Mongodin EF, Fricke WF, Gajer P. et al. The pangenome structure of Escherichia coli: comparative genomic analysis of E. coli commensal and pathogenic isolates. J Bacteriol. 2008;190(20):6881-93
Author contact
Corresponding author: Tel.: + 55 71 3283-8940. E-mail: asilvabr (A. Silva).

 Global reach, higher impact
Global reach, higher impact