ISSN: 1839-9940
J Genomics 2016; 4:29-41. doi:10.7150/jgen.16885 This volume Cite
Research Paper
De novo Assembly and Analysis of the Chilean Pencil Catfish Trichomycterus areolatus Transcriptome
1. Department of Biology, University of Nebraska at Omaha, Omaha, Nebraska 68182, USA;
2. Center for Environmental Health and Toxicology, University of Nebraska Medical Center, Omaha, Nebraska 68198, USA;
3. Department of Environmental, Agricultural and Occupational Health, University of Nebraska - Medical Center, Omaha, NE, 68198-6805, United States;
4. Departamento de Biología, Universidad de La Serena, La Serena, Chile;
5. Centro de Estudios Avanzados en Zonas Aridas, La Serena, Chile;
6. Instituto de Investigación Multidisciplinar en Ciencia y Tecnología, Universidad de La Serena, La Serena, Chile.
Published 2016-9-6
Abstract
Trichomycterus areolatus is an endemic species of pencil catfish that inhabits the riffles and rapids of many freshwater ecosystems of Chile. Despite its unique adaptation to Chile's high gradient watersheds and therefore potential application in the investigation of ecosystem integrity and environmental contamination, relatively little is known regarding the molecular biology of this environmental sentinel. Here, we detail the assembly of the Trichomycterus areolatus transcriptome, a molecular resource for the study of this organism and its molecular response to the environment. RNA-Seq reads were obtained by next-generation sequencing with an Illumina® platform and processed using PRINSEQ. The transcriptome assembly was performed using TRINITY assembler. Transcriptome validation was performed by functional characterization with KOG, KEGG, and GO analyses. Additionally, differential expression analysis highlights sex-specific expression patterns, and a list of endocrine and oxidative stress related transcripts are included.
Keywords: de novo transcriptome, assembly, catfish, Trichomycterus areolatus
Introduction
Trichomycterus areolatus (described by Valenciennes, 1846) is a threatened pencil catfish found in Chile [1, 2]. This freshwater stream fish has a broad distribution ranging from the Coquimbo to the Los Lago regions of Chile (30° S and 43° S respectively) [1, 3]. The genus Trichomycterus is found throughout freshwater ecosystems in both Central and South America and includes over 120 species [4]. Individual species are commonly restricted to specific river areas either by geographical restriction or habitat preference [5]. As a result, T. areolatus displays an intricate integration with its environment; yet the species is fairly distributed throughout the region, providing an opportunity for its use as a model sentinel organism to study environmental stress in Chile's freshwater streams [6,7]. As a benthic fish, T. areolatus displays interactions with riverine sediments and substrates [8]. This species tends toward a generalist eating strategy, eating aquatic insects [9] as well as the surface organisms present on stream, plant, and rock surfaces [10]; consequently, its food preferences are known to vary by season [4].
The Trichomycterus fish generally range between 50 - 150 mm [11]. The mature fish in this species will reach average sizes of 56 and 51mm for males and females respectively [12]. They display poor secondary sex characteristics, making the identification of sex by external means difficult [13]. The spawning season takes place during October and November, based on observation of the female gonads rapidly increasing in size just before the season begins [13]. The females are capable of releasing eggs several times throughout the spawning season. The males also exhibit numerous fertilization events throughout the spawning season; however, the male fish in this species do not participate in progeny care [14]. The T. areolatus genome is described as diploid, with the typical individual displaying 2n = 54 chromosomes; however, intra-individual variation has been reported [15]. This chromosome number appears common among the Trichomycterus genus and suggests conservation.
Genomes and/or transcriptomes have only been developed for fish that have either economic or scientific value including: Salmo salar (Atlantic salmon), Cyprinus carpio (common carp), Pimephales promelas (fathead minnow), and Danio rerio (zebrafish). There are limited transcriptomic resources available for fish native to South America, and none available of the Trichomycterids. Current molecular research with this organism is limited, and includes a karyotype [15], seasonal variation in biomarkers [8], local industrial discharge impacts [6], agricultural disturbances [16], microsatellite loci for conservation resources [17], and population genetics [18].
In this paper, we detail the assembly and analysis of the transcriptome for T. areolatus. Fish samples obtained from Chile were sequenced by next-generation sequencing and later assembled into a de novo transcriptome using Trinity. Transcriptome validation analyses are included and discussed. Transcriptomic data is deposited at NCBI SRA under accession SRP077018 and the completed assembly is freely available.
Results and Discussion
Transcriptome Characteristics
The constructed transcriptome assemblies are available at http://www.davislab.net/trichomycterus/. Assemblies available include: a complete transcriptome nucleotide assembly, a representative assembly (see Methods) consisting of 64,385 unique transcripts, and a translated protein assembly of the representative transcripts. De novo assembly was accomplished using TRINITY. Two samples (whole male and whole female adult fish) provided RNA for the assembled transcriptome, which was assembled from Illumina® paired-end reads. A resulting 41.8Gb of output (Table 1) was utilized to create the assembly. General statistics of the assembled representative transcriptome are included in Table 2.
Transcriptome Tissue Sequencing Details. RNA samples were from 4.4 and 5.4ug of RNA for male and female fish, respectively. Sequencing was performed on an Illumina® Hi-Seq 2500. BP: base pair, GC: G-C nucleotide ratio.
| Tissue | Total Reads | Total Output (bp) | GC Content (%) |
|---|---|---|---|
| Whole Female | 328,721,780 | 32,872,178,000 | 48% |
| Whole Male | 88,794,542 | 8,879,454,200 | 47% |
Trichomycterus areolatus Representative Transcriptome Characteristics. The representative Trichomycterus areolatus transcriptome assembly was analyzed for general characteristics listed above. Putative protein coding transcripts were included and identified by TransDecoder. Redundant transcripts were removed by CD-HIT which collapses redundant and highly similar sequences into consensus sequences.
| Total Transcripts | Mean Length (bp) | Median Length (bp) | N50 | GC Content |
|---|---|---|---|---|
| 64889 | 1484.85 | 857 | 2671 | 47.5% |
Organism Phylogenetics
Assembled T. areolatus sequences were selected and aligned to publically available sequences from various fish to examine phylogenetic relatedness by identifying orthologs (Table 3). Of the fish compared, Ictalurus punctatus (Channel catfish) showed the highest relationship (87.6%) in a concatenated set of conserved sequences, followed closely by Cyprinus carpio (Common carp) at 86.5%. Intra-genus phylogenetic comparison of Trichomycterus was not possible due to a lack of published sequences.
Phylogenetic Comparison to Other Fish. Transcripts produced in this study were concatenated and aligned to published sequences of fish species and a percent identity matrix was computed. This analysis utilized Trichomycterus areolatus transcripts: Ta_155828, Ta_53325, Ta_192266, Ta_56196, and Ta_56194 for cytochrome c oxidase subunit III, HSP70, NADH dehydrogenase subunit 5, estrogen receptor, and glutathione s-transferase kappa 1, respectively.
| Species | Percent Identity |
|---|---|
| Ictalurus punctatus (Channel catfish) | 87.6 |
| Cyprinus carpio (Common carp) | 86.5 |
| Pimephales promelas (Fathead minnow) | 84.5 |
| Oncorhynchus mykiss (Rainbow trout) | 82.5 |
| Salmo salar (Salmon) | 82.5 |
| Carassius auratus (Gold fish) | 76.4 |
| Danio rerio (Zebra fish) | 67.1 |
Assessing Full-length Transcript Coverage
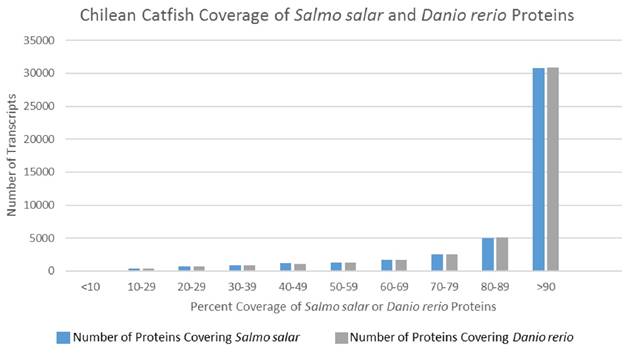
In an effort to deduce the full-length nature of the transcripts within the representative assembled transcriptome, coverage histograms arising from alignments with non-redundant protein sequences of two model organisms—Salmo salar and Danio rerio—were produced. Model organisms were selected over more closely related organisms (Table 3) due to their more complete publically available transcriptomes. Figure 3 illustrates a histogram for the distribution of transcript length when the representative T. areolatus transcriptome was compared to related species. Length coverage exceeding 90% of the other organism's transcript length was found in 64.7% of T. areolatus protein sequences upon alignment with Salmo salar proteins and 69.1% of Danio rerio proteins (Figure 3). Accounting for genetic differences between these organisms, these data suggest that the produced representative transcriptome has a high degree of full-length transcripts.
Transcript Coverage of Two Model Organisms. Coverage of Salmo salar and Danio rerio predicted proteins by Trichomycterus areolatus predicted proteins. Predicted polypeptide sequences produced in this study were BLASTed against publically available non-redundant Salmo salar proteins (count = 112,089) and Danio rerio proteins (count = 81,931). The length of the local alignment region reported by the BLASTp algorithm was subsequently divided by the length of the query sequence. Compilation of these results indicated that a vast majority of Trichomycterus areolatus predicted protein sequences exhibited greater than 90% coverage of both Danio rerio (64.7%) and Salmo salar (68.9%) protein sequences, suggesting that the assembly produced a high degree of full-length transcripts.
Functional Analyses - Putative Transcript Functional Characterization
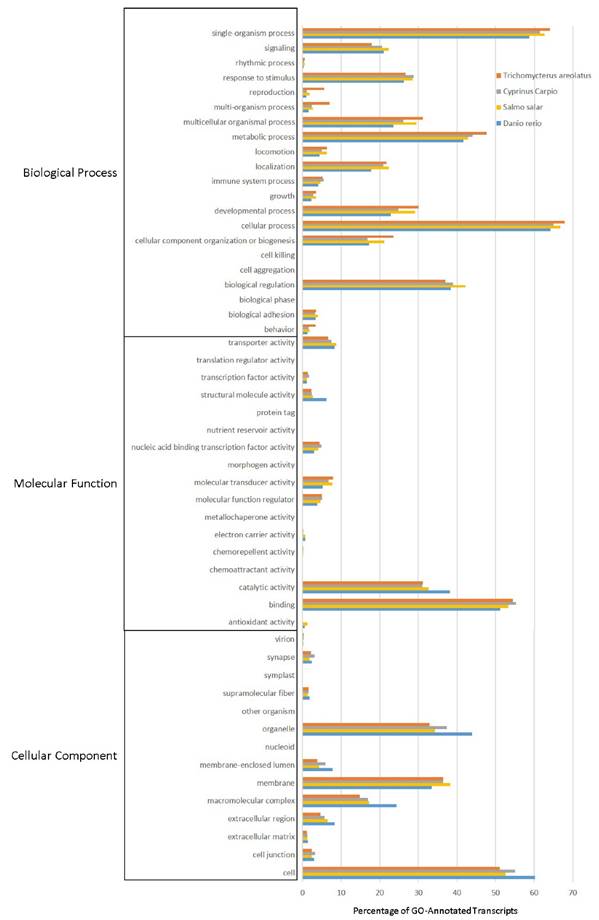
Multiple analyses exist that allow transcripts to be annotated and grouped by function. This includes Gene Ontology (GO), Eukaryotic Orthologous Groups (KOG), and Kyoto Encyclopedia of Genes and Genomes (KEGG) analyses. These annotation and grouping algorithms were applied to examine the putative function of transcripts, and as a quality control technique to evaluate transcriptome completeness when compared to well-developed transcriptomes from Danio rerio, Salmo salar, and Cyprinus carpio. Figure 4 details the GO analysis, which is capable of classifying predicted gene products by cellular component, molecular function, and biological process [19]. GO groupings between organisms suggest that the selected fish all share a pattern of functional distribution of gene products.
Gene Ontology (GO) Analysis of the Trichomycterus areolatus Transcriptome. GO functional analysis was performed on assigned proteins in order to evaluate transcript function and the overall completeness of the isolated transcriptome. GO terms were given for each of the T. areolatus predicted proteins as well as the proteomes of Salmo salar, Cyprinus carpio, and Danio rerio (retrieved from NCBI). The distribution of protein functions closely match one another, suggesting the assembled transcriptome is complete.
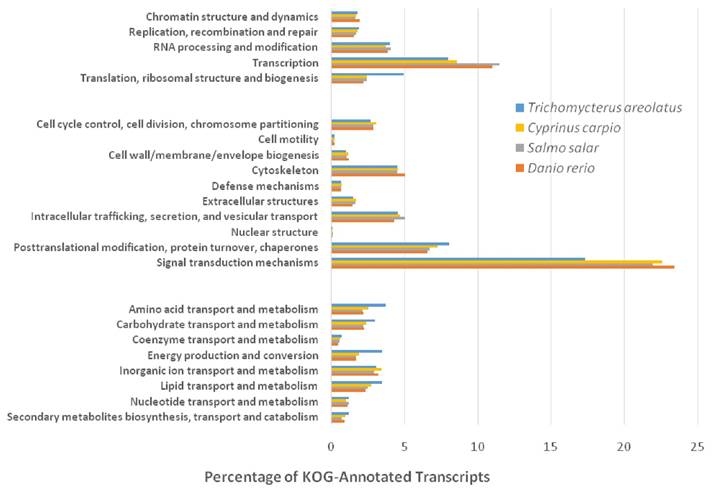
Similarly, Figure 5 details the KOG transcriptomic analysis; this database allows the user to compare against a collection of seven eukaryote genomes of a diverse set (e.g. human, fly, parasite, plant, worm, fungi) of model organisms [20]. These annotated proteins are organized by function into clusters of eukaryotic orthologous groups. Orthologs typically have similar functions among different organisms and serve as an effective means of identifying putative functions of gene products [20]. The KOG analysis of T. areolatus resulted in KOG groups classifying 81% of transcripts. Finally, KEGG analysis was performed to identify biologically relevant pathways of function for predicted protein products from the representative transcriptome [21]. T. areolatus was compared to Danio rerio, Salmo salar and Cyprinus carpio as is shown in Figure 6. Distribution of group numbers also appears consistent as related fish were evaluated. Overall, each annotation and grouping algorithm showed close association with T. areolatus and the related interrogated species, providing evidence of the completeness and consistency of the transcriptome.
Eukaryotic Orthologous Groups (KOG) Characterization of Trichomycterus areolatus Transcripts. Putative transcript functions were assessed and transcriptome completeness was evaluated using KOG analysis. The Trichomycterus areolatus transcriptome and mRNA nucleotide entries from NCBI of Cyprinus carpio, Salmo salar, and Danio rerio were assigned KOG terms. The three transcriptomes have similar distributions, supporting the completeness of the Trichomycterus areolatus transcriptome.
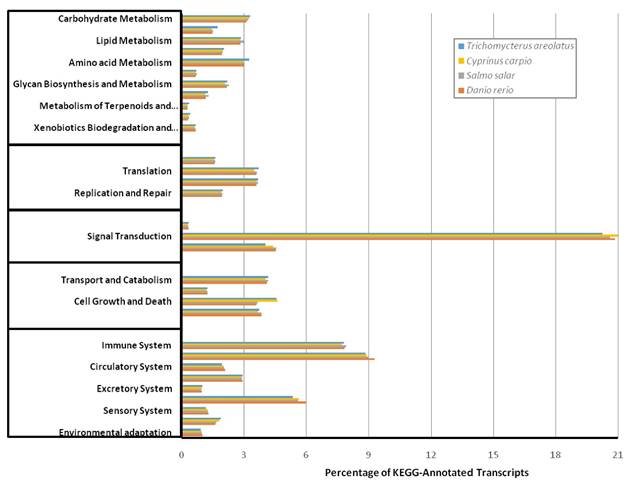
Kyoto Encyclopedia of Genes and Genomes (KEGG) Transcriptomic Analysis. KEGG analysis was performed to functionally describe transcript functions and evaluate transcriptome completeness. To serve as comparisons mRNA sequences for Danio rerio, Salmo salar, and Cyprinus carpio were retrieved from NCBI and characterized into KEGG pathways. The percent distribution shows a similar proportion among compared species indicating a complete transcriptome for Trichomycterus areolatus.
Differential Expression Analysis
Table 4 details unique transcripts which were differentially regulated between male and female (sex determined by necropsy) T. areolatus samples. Non-inherited transcripts that are unique to individual organisms (e.g. MHC molecules via rearrangement) were excluded.
Differentially Expressed Transcripts. The resultant male and female Trichomycterus areolatus sequence files were interrogated to assess relative differential transcript expression. Unique transcripts, demonstrating changes greater than or equal to 10-fold, are identified by homology (if available) to known proteins, and only the most differentially expressed isoform is presented. Non-inherited transcripts that are unique to individual organisms (e.g. MHC molecules via rearrangement or similar immune transcripts) were excluded. Notably, many transcriptional differences are related to sex-specific expression.
| Transcriptomic ID | Fold Change | Name |
|---|---|---|
| Male differentially upregulated transcripts | ||
| TRICH01_163992 | 15 | parvalbumin beta-1 |
| TRICH01_133698 | 12 | hemoglobin subunit beta-2 |
| TRICH01_83415 | 11 | sperm acrosome membrane-associated protein 4 |
| TRICH01_57563 | 10 | endonuclease domain-containing 1 protein |
| Female differentially upregulated transcripts | ||
| TRICH01_101761 | 38604 | microsomal triglyceride transfer protein large subunit |
| TRICH01_222696 | 96 | complement C4 |
| TRICH01_180471 | 96 | alpha-2-macroglobulin |
| TRICH01_143497 | 56 | 3-hydroxyacyl-CoA dehydrogenase type-2 |
| TRICH01_54871 | 42 | vitellogenin 4 |
| TRICH01_100604 | 38 | pyruvate dehydrogenase phosphatase regulatory subunit |
| TRICH01_142598 | 35 | CD59 |
| TRICH01_51308 | 29 | ribonucleoside-diphosphate reductase subunit M2 |
| TRICH01_208172 | 22 | coiled-coil domain-containing protein 36 |
| TRICH01_51310 | 21 | Jouberin |
| TRICH01_100599 | 19 | PEX5 |
| TRICH01_100601 | 17 | histone-lysine N-methyltransferase ASH1L |
| TRICH01_100600 | 16 | A-kinase anchor protein |
Among the most highly differentially regulated transcripts, the gene product of A2-macroglobulin (A2M) functions as a protease inhibitor and binds growth factors [22]. A2M, increased in the female by 96-fold, has been shown to increase endogenous production of estradiol resulting in increased follicular cell proliferation and oocyte maturation [22].
Vitellogenin is an egg yolk lipoprotein precursor that serves as an essential material for oocyte development [23, 24, 25]. Vitellogenin production is under estrogenic control, and was upregulated in the female T. areolatus 42 fold or more when compared to the male. The female liver (the primary vitellogenin production site) secretes the lipoprotein, which is then transported from the systemic circulation into oocytes within the developing ovary. Because vitellogenins are large lipoproteins, mobilization requires the microsomal triglyceride transfer protein for delivery to the oocyte. Microsomal triglyceride transfer protein is crucial in final yolk lipoprotein assembly [26], and is seen upregulated in the female by 38,604-fold.
Throughout the process of vitellogenesis, the female organism undergoes metabolic changes, shifting energy usage, in order to support the growth of newly forming oocytes. Moreover, lipid metabolism is essential during ovarian development, including lipid storage, oxidation, and as previously mentioned, mobilization. Under some conditions such as starvation, the female may utilize stored lipids for transfer to the growing oocyte, or subsequent oxidation to meet dietary needs when intake is low [27]. The increased expression (56-fold) of 3-hydoxyacyl-CoA dehydrogenase suggests increased lipid breakdown and usage in the female most likely stimulated by increased energy requirements during vitellogenesis.
Similarly, the transcriptomic profile of the female demonstrates increased glycolytic activity, likely also to support the energy-intensive vitellogenesis. The pyruvate dehydrogenase complex is a highly regulated system that connects glycolysis with the TCA cycle. Activation of the complex is enhanced by pyruvate dehydrogenase phosphatase (PDP) [28]. The observed 38-fold increased expression of pyruvate dehydrogenase phosphatase would promote pyruvate progression through the TCA cycle and yield additional cellular energy in the form of ATP.
The complement system in fish is a primary component of their innate immune system and is suggested to be more beneficial to organism defense than that of mammalian systems [29]. Complement proteins are produced by the liver in an inactive form, where they are activated by proteolysis, ultimately leading to either opsonization, phagocytosis, or lysis of the pathogen. Vertebrate fish have multiple isoforms and isotypes of complement proteins, enabling the organism's immune system to recognize a wide range of pathogens, which is crucial due to slow lymphocyte proliferation, and limited antibody production and affinity in the fish immune system [30,31]. Notably, catfish as well as other teleost species, practice external fertilization which may expose the developing embryo and developing fish to waterborne threats, including pathogens. Maternal complement proteins are transferred from the female to the egg and serve to protect the embryos until the immune system and lymphoid organs are competent enough to protect the developing fish [32]. Thus, it is expected that maternal complement proteins, especially CD59 which acts as a stabilizer to prevent premature complement activation, would be upregulated. Accordingly, the CD59 gene is upregulated by 35-fold in the female. However, seasonal and temperature variations cause complement proteins to vary in expression, accounting for some elevated complement proteins in the male [33].
Relative increases in male-associated transcripts are also presented in Table 4. Parvalbumins are calcium binding proteins found in white, fast twitch skeletal muscle of most fish species. In muscular tissues it acts to sequester calcium, accelerating muscle relaxation. High expression of parvalbumins can promote quicker muscle relaxation, however overexpression leads to smaller mitochondrial densities in slow twitch muscles [34]. More recently, the beta-1 parvalbumin isoform in carp seminal plasma has been characterized [35], and while the specific details of its function remain undefined, it is thought to play a role in sperm motility. Studies in carp have shown that sperm are not mobile without calcium, and initiation of motility is seen with an increase in intracellular calcium levels. The influx of calcium has been suggested to be the initiating factor of sperm motility, and therefore, the presence of parvalbumin as a calcium binding protein may play an essential role in fish sperm function. Additionally, studies in mammalian organisms have suggested parvalbumin regulation in calcium-mediated spermatogenesis and testosterone production [36]. Accordingly, the male transcriptome demonstrates a 15-fold increase in beta-1 parvalbumin expression.
Though many studies do not identify significant sex differences of hemoglobin levels in fish [37], Falahatkar et al. [38] discovered decreased hemoglobin concentrations and hematological changes in juvenile Acipenser stellatus that were treated with estradiol for 7-9 months. They predict estradiol may have an inhibitory effect on erythropoiesis (the production of red blood cells). Hemoglobin transcription levels in the male were found to be 12-fold upregulated compared to the female.
During mammalian fertilization, the acrosome surrounding the sperm head releases acrosomal hydrolases that enables the sperm and egg to combine [39]. The acrosomal-egg interaction, and release of hydrolytic enzymes, are thought to be stabilized by the outer acrosomal membrane matrix interactions, and the specificity of the binding interactions are essential in release of hydrolases [39]. The sperm acrosome membrane associated protein 4 is retained following the egg-sperm binding, localizing to the inner acrosomal membrane in human sperm, and may play a role in fertilization. Of note, teleost organisms differ in fertilization strategies: their sperm do not possess outer acrosomal membranes and therefore, do not undergo the same acrosomal reaction seen in mammals. Interestingly, recent catfish transcriptome studies have identified a homolog of sperm acrosome membrane-associated protein 4 [40], though the specific function in catfish has not been elucidated [41]. A homolog to acrosome membrane associated protein 4 was observed in this male, expressed at 11-fold higher compared to the female.
Consideration as a Sentinel Organism
This transcriptome provides a resource to utilize T. areolatus as a sentinel organism or a “canary in the coal mine” for biological effects that may be experienced by local wildlife and nearby human populations. In North America, gene expression biomarkers have been readily applied to environmental monitoring for anthropogenic pollutants [42,43,44,45]. For example, fathead minnows are commonly used environmental sentinel in studies on agricultural runoff [46,47,48], waste water treatment plant effluent [49,50] and industrial waste effluent [51]. The growing reliance on transcriptomic tools in the field of environmental toxicology has been due to their increasing availability for non-model organisms as well as the mechanistic insight they provide for prediction of adverse outcomes at the whole organismal and possibly population level [52,53].
Chile is experiencing significant economic growth driven by agricultural and industrial development which puts considerable pressure on the freshwater resources across much of the country [54,55]. However, scientifically documented adverse impacts on water quality are largely absent, particularly with respect to biologic impacts on sentinel species. Thus, a need exists to develop environmental sentinel organisms suitable for study across the country. T. areolatus is an ideal candidate, as it can be found across much of the range of many rivers in Chile from the mountain watersheds to the low elevation, coastal regions of the country. Furthermore, the fish can be found associated with the benthos in streams where the water is merely a few centimeters in depth. Finally, the previously mentioned benthic behavior of the fish causes it to be intimately associated with sediments, therefore it is susceptible to contaminants that are affiliated with the water and sediment [56]. To provide a resource for the study of environmental systems associated with this organism, specific genes were identified for convenience (Table 5). These selected genes of interest will serve as markers of reproductive dysfunction and/or oxidative stress, in an upcoming manuscript currently in progress.
Trichomycterus areolatus Environmental Sentinel Biomarkers. Genes linked to endocrine disruption and/or oxidative stress were identified within the transcriptome assembly for convenience in developing Trichomycterus areolatus as an environmental sentinel organism. Danio rerio sequences were used as queries to BLAST the full assembly to identify putative homologs. Protein isoforms were differentiated based on query sequence annotation and bitscore.
| Gene Name | Gene Symbol | Transcriptomic ID | Transcript Length (bp) | Query ID | Bit Score |
|---|---|---|---|---|---|
| Androgen Receptor | AR | TRICH01_58265 | 4455 | NP_001076592.1 | 788 |
| Aromatase | CYP19a1 | TRICH01_14384 | 284 | AAB65788.1 | 759 |
| Aryl Hydrocarbon Receptor | AHR | TRICH01_166983 | 3366 | NP_001019987.1 | 593 |
| Aryl Hydrocarbon Receptor 2 | AHR2 | TRICH01_225654 | 2827 | NP_571339.1 | 905 |
| Cytochrome P450 1A1 | CYP1a1 | TRICH01_117246 | 2028 | NP_571954.1 | 820 |
| Estrogen Receptor Alpha | ESRa | TRICH01_56196 | 4380 | AAK16740.1 | 729 |
| Estrogen Receptor Beta 1 | ESRb1 | TRICH01_211240 | 4700 | CAC93848.1 | 673 |
| Estrogen Receptor Beta 2 | ESRb2 | TRICH01_95151 | 3578 | CAC93849.1 | 796 |
| Follicle Stimulating Receptor | FSHR | TRICH01_111037 | 3596 | AAP33512.1 | 996 |
| Forkhead Box L2 | FOXL2 | TRICH01_143206 | 1866 | AAI16586.1 | 370 |
| Heat Shock Protein 70 | HSP70 | TRICH01_53325 | 2625 | AAF70445.1 | 1216 |
| Heat Shock Protein 90 Alpha 1 | HSP90a1 | TRICH01_121146 | 2863 | NP_571403.1 | 1292 |
| Heat Shock Protein 90 Alpha 2 | HSP90a2 | TRICH01_121144 | 2926 | AAI63166.1 | 1278 |
| Metallothionein | MT | TRICH01_130427 | 554 | AAS00513.1 | 53 |
| Superoxide Dismutase | SOD | TRICH01_28552 | 2500 | NP_571369.1 | 261 |
| Thyroid Receptor Alpha | THRa | TRICH01_196629 | 2500 | AAA99811.1 | 760 |
| Thyroid Receptor Beta | THRb | TRICH01_20904 | 2406 | AF109732_1 | 732 |
| Vitellogenin 1 | VTG1 | TRICH01_101739 | 2851 | AF406784_1 | 1224 |
The availability of a complete transcriptome for T. areolatus provides a valuable resource relative to the development of this species as a sentinel organism. The unique niche that T. areolatus fills in Chile will allow it to be useful when considering water and sediment contamination.
Methods
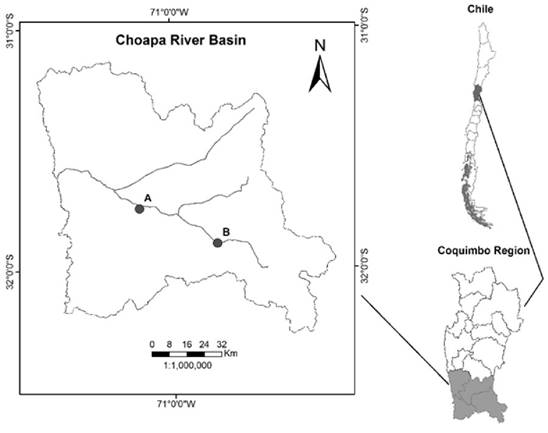
Tissue Collection and RNA Preparation: Whole fish were collected from the Choapa River basin in the Coquimbo region (region VII) of Chile in July 2015 under fishing authorization #2017 by the Chilean Subsecretary of Fisheries and Aquaculture. Specifically, a whole male and female with clear sexual differentiation were sampled from a downstream and upstream site respectively. The river sampling site coordinates are shown in Figure 2. This watershed is used intensively for agricultural production.
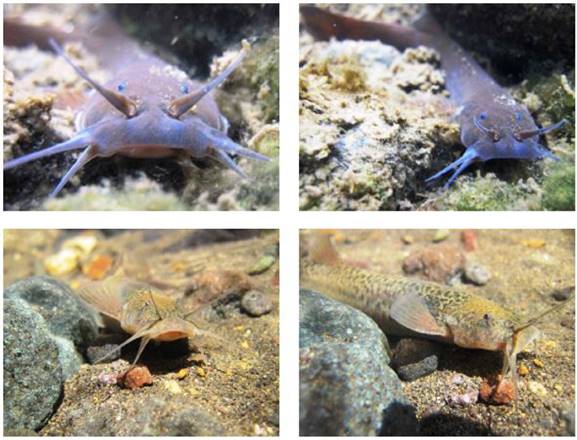
Representative Photo of Organism-Trichomycterus areolatus organism photos of unspecified sex taken from two different locations in Chile, the Maule River (top) and Pangue River (bottom). Trichomycterus areolatus displays intimate contact with both sediment and the water column. Morphology and behavior is typical of fish who dwell in fast-moving streams. Photos were provided by courtesy of Pablo Reyes, Fundación Ictiológica (Chile) [70].

Choapa River Basin Tissue Sampling Sites-The male and female fish samples were collected from the downstream site “A” (altitude 243m, Lat; Lon: -31.749639; -71.160722) and upstream site “B” (altitude 792m, Lat; Lon: -31.89675; -70.783056) respectively. This river basin is proximate to heavy agricultural practice and downstream of heavy metals mining (e.g. copper). The stream itself is predominately supplied by glacial melt from the Andean mountains. Historically, this system drained into the Pacific Ocean but is becoming an isolated system due to the deleterious effects of climate change.
Fish samples were prepared and immediately submerged in RNAlater® (Ambion) according to specific manufacturer recommendations to preserve RNA integrity. The samples were mechanically homogenized in the presence of Qiagen Lysis Buffer RLT; immediately following, a Qiagen RNeasy Mini Plus isolation kit was used to isolate and purify organism total RNA. The resultant RNA was quantified by Thermo Scientific™ NanoDrop 2000c and verified for integrity with a bleach denaturing agarose electrophoresis gel [57]. Prior to sequencing, the purified RNA was stored at -80° Celsius with minimal handling and freeze-thawing cycles.
High-throughput sequencing: Prior to sequencing, a TruSeq® RNA Sample Preparation Kit was used to prepare the library for sequencing. An Illumina® HiSeq2500 next generation sequencer was used to generate paired-end 101bp reads. Sequencing was performed at the University of Nebraska Medical Center sequencing core. The whole female and male tissue reads were used exclusively in transcriptome construction.
Data Processing and Assembly: The resulting sequence reads were first processed with FastQC [58] to evaluate sequence quality. Next, PRINSEQ was used to both trim reads by quality score, and apply a dusting technique [59]. Transcriptome assembly was performed de novo with TRINITY assembler [60, 61, 62]. The resultant initial transcriptome assembly was searched for predicted coding sequences at least 50 amino acids long using TransDecoder [61], BLASTed against NCBI Refseq metazoan protein sequences from March 2016 (retained if the bit score was 50 or higher), and filtered for microbial sequence contamination using the same BLAST [63] method. Noncoding RNA was removed based on homology to Rfam sequences of Danio rerio, and highly similar sequences were collapsed using CD-HIT to form the “representative transcriptome”. This Transcriptome Shotgun Assembly project has been deposited at DDBJ/ENA/GenBank under the accession GEVC00000000. The version described in this paper is the first version, GEVC01000000.
Percent Identity Matrix: Sequences acquired from the current assembly were aligned with sequences of previously published sequences from different species to evaluate phylogenic relationships within fish. Alignments were trimmed with Gblocks [64, 65], concatenated manually into a single ordered sequence, and aligned with ClustalW [66] to produce a percent identity matrix.
Transcript Coverage against Model Organisms: A FASTA file was compiled containing all protein sequences derived from the T. areolatus transcriptome; the sequences were BLASTed against all non-redundant Salmo salar and Danio rerio protein sequences retrieved from NCBI. The BLASTp algorithm was used to establish local alignments with E-values smaller than 1e-5. The sequences demonstrating the highest-scored alignments were kept. The length of each high-scoring alignment was subsequently compared to the overall length of the reference sequence to obtain the coverage, after which a count of the unique sequences present in a particular range of coverage was obtained through the Linux command line. The results were then compiled into Figure 3.
Gene Ontology: BLASTp was used to assign the top hit for the TransDecoded T. areolatus proteins and the proteomes of Salmo salar, Cyprinus carpio, and Danio rerio (retrieved from NCBI January 29th, 2016) against the NCBI non-redundant database (GI list: Arabidopsis thaliana, Caenorhabditis elegans, Danio rerio, Dictyostelium discoideum, Drosophila melanogaster, Escherichia coli, Gallus gallus, Homo sapiens, Mus musculus, Rattus norvegicus, Saccharomyces cerevisiae, and Schizosaccharomyces pombe retrieved June 10th, 2016). The resulting file was scanned by BLAST2GO (version 2.8.0) [67] with the b2g_feb16 GO database; once terms were assigned, level 2 GO ID terms were utilized for a comparison with all groups.
KOG: Analysis included translating publically available transcripts from chosen fish species using TransDecoder version 3.0 [61]. These results were aligned using RPS-BLAST (E-value of ≤ 1e-5) to the NCBI KOG database (version 3.14).
KEGG: The KEGG analysis was conducted by uploading organism transcriptomic sequences to the KEGG Automatic Annotation Server (KAAS) where they are processed by BLAST and GHOST comparisons against a database of KEGG genes. Ortholog assignments were returned and graphed to illustrate a transcriptome pathway comparison.
Differential Expression Analysis: Differential transcript expression analysis of T. areolatus transcript files were aligned to the assembled reference sequences. Reads were mapped with Bowtie [68], and RSEM [69] was used to quantify differential expression values. Transcripts Per Kilobase Million (TPM) values were calculated to produce fold changes. Transcripts were then annotated by homology to NCBI Refseq proteins.
Genes of Interest: To identify genes related to endocrine and/or oxidative stress, query protein sequences from published sequences available on NCBI were obtained. Sequences from Danio rerio were used with BLASTp to find top hits in the putative coding sequence transcriptome (T. areolatus). Top hits were reviewed for the highest bit score (in combination with E-value) and included in Table 5. Protein isoforms were differentiated based off of chosen model organism annotation and could vary between query species (e.g. estrogen receptor alpha, beta).
Acknowledgements
This work was supported by NIH grant P20GM103427 (P.H.D.). The University of Nebraska Medical Center Genome Sequencing Center provided RNA sequencing, and is additionally supported by NIH grant 1P30GM110768 and The Fred & Pamela Buffett Cancer Center Support Grant P30CA036727. Additional financial support was provided by the Fund for Undergraduate Scholarly Experiences (FUSE) and the Biomedical Research Training Program at the University of Nebraska. Jonathan Ali was partially supported by Robert G. Daugherty Water for Food Institute Graduate Assistantship, University of Nebraska, and the Center for Global Health and Development 2015 Summer Research Fellowship Program, University of Nebraska Medical Center. This publication's contents are the sole responsibility of the authors and do not necessarily represent the official views of the aforementioned funding agencies.
Competing Interests
All investigators have declared that no competing interests existed.
References
1. Arratia G, Rojas G, Chang A. Géneros de peces de aguascontinentales de Chile. Museo Nacional de Historia Natural PublicacionOcas. 1981;34:3-108
2. Arratia G. Preferencias de habitat de pecessiluriformes de aguascontinentales de Chile (Fam. Diplomystidae y Trichomycteridae). Studies On Neotropical Fauna and Environment. 1983;18(4):217-237
3. Dyer BS. Revisiónsistemáticabiogeográfi ca de lospecesdulceacuícolas de Chile. EstudiosOceanologicos. 2000;19:77-98
4. Sergio S, Rodrigo P, Vila I. Trophic niche overlap between two Chilean endemic species of Trichomycterus (Teleostei: Siluriformes). RevistaChilena de Historia Natural. 2007;80(4):431-437
5. Lima SM, Costa W. Trichomycterus giganteus (Siluriformes: Loricarioidea: Trichomycteridae): a new catfish from the Rio Guandu basin, southeastern Brazil. Zootaxa. 2004;761:1-6
6. Chiang G, McMaster ME, Urrutia R, Saavedra MF, Gavilán J, Tucca F, Munkittrick KR. Health status of native fish (Perciliagillissi and Trichomycterus areolatus) downstream of the discharge of effluent from a tertiary-treated elemental chlorine-free pulp mill in Chile. Environmental Toxicology and Chemistry. 2011;30(8):1793-1809
7. Arciszewski TJ, Munkittrick KR. Development of an adaptive monitoring framework for long-term programs: An example using indicators of fish health. Integrated Environmental Assessment and Management. 2015;11(4):701-718
8. Chiang G, Munkittrick KR, Urrutia R, Concha C, Rivas M, Diaz-Jaramillo M, Barra R. Liver ethoxyresorufin-O-deethylase and brain acetylcholinesterase in two freshwater fish species of South America; the effects of seasonal variability on study design for biomonitoring. Ecotoxicology and environmental safety. 2012;86:147-155
9. Habit E, Victoriano P, Campos H. Ecologíatrófica y aspectosreproductivos de Trichomycterus areolatus (Pisces, Trichomycteridae) enambienteslóticosartificiales. Revista de Biología Tropical. 2005;53:195-210
10. Aranha JMR, Takeuti DF, Yoshimura TM. Habitat use and food partitioning of the fishes in a coastal stream of Atlantic forest, Brazil. Revista de Biología Tropical. 1998;46:951-959
11. Alencar A, Costa W. Trichomycterus pauciradiatus, a new catfish species from the upper rio Paraná basin, southeastern Brazil (Siluriformes: Trichomycteridae). Zootaxa. 2006;1269:43-49
12. Manríquez A, Huaquín L, Arellano M, Arratia G.Aspectosreproductivos de Trichomycterus areolatusValenciennes, 1846 (Pisces. Teleostei: Siluriformes) enrío Angostura, Chile. Stud Neotrop Fauna Environ. 1988;23(2):89-102
13. Chiang G, Munkittrick KR, Saavedra MF, Tucca F, McMaster ME, Urrutia R, Tetreault G, Barra R. Seasonal changes in reproductive endpoints in Trichomycterus areolatus (Siluriformes: Trichomycteridae) and Perciliagillissi (Perciformes, Perciliidae), and the consequences for environmental monitoring. Studies on Neotropical Fauna and Environment. 2011;46(3):185-196
14. Pavlov DA, Emel´yanova NG, Novikov GG. Reproductive Dynamics. In: (ed.) Jakobsen T, Fogarty MJ, Megrey BA, Moksness E. Fish Reproductive Biology, Implications for Assessment and Management. Chichester, UK: Wiley-Blackwell publishing. 2009:48-90
15. Colihueque N, Corrales O, Parraguez M. Karyotype and nuclear DNA content of Trichomycterus areolatus (Siluriformes, Trichomycteridae). Genetics and Molecular Biology. 2006;29(2):278-282
16. Orrego R, Adams SM, Barra R, Chiang G, Gavilan JF. Patterns of fish community composition along a river affected by agricultural and urban disturbance in south-central Chile. Hydrobiologia. 2009;620(1):35-46
17. Muñoz-Rojas P, Quezada-Romegialli C, Véliz D. Isolation and characterization of ten microsatellite loci in the catfish Trichomycterus areolatus (Siluriformes: Trichomycteridae), with cross-amplification in seven Trichomycterinae species. Conservation Genetics Resources. 2012;4(2):443-445
18. Quezada-Romegialli C, Fuentes M, Véliz D. Comparative population genetics of Basilichthysmicrolepidotus (Atheriniformes: Atherinopsidae) and Trichomycterus areolatus (Siluriformes: Trichomycteridae) in north central Chile. Environmental Biology of Fishes. 2010;89(2):173-186
19. Gene Ontology Consortium. The Gene Ontology project in 2008. Nucleic Acids Res. 2008Jan;36:D440-4 Epub 2007 Nov 4
20. Tatusov RL, Fedorova ND, Jackson JD, Jacobs AR, Kiryutin B, Koonin EV, Krylov DM, Mazumder R, Mekhedov SL, Nikolskaya AN, Rao BS, Smirnov S, Sverdlov AV, Vasudevan S, Wolf YI, Yin JJ, Natale DA. The COG database: an updated version includes eukaryotes. BMC Bioinformatics. 2003Sep11;4:41. Epub 2003 Sep 11
21. Kanehisa M, Goto S. KEGG: kyoto encyclopedia of genes and genomes. Nucleic Acids Res. 2000Jan1;28(1):27-30
22. Ireland JLH, Jimenez-Krassel F, Winn ME, Burns DS, Ireland JJ. Evidence for autocrine or paracrine roles of α2-macroglobulin in regulation of estradiol production by granulosa cells and development of dominant follicles. Endocrinology. 2004;145(6):2784-2794
23. Lim EH, Ding JL, Lam TJ. Estradiol-induced vitellogenin gene expression in a teleost fish, oreochromis aureus. General and Comparative Endocrinology. 1991;82(2):206-214
24. Lattier DL, Gordon DA, Burks DJ, Toth GP.Vitellogenin gene transcription. A relative quantitative exposure indicator of environmental estrogens. Environmental Toxicology. 2001;20(9):1979-1985
25. García-Reyero N, Raldúa D, Quirós L. et al. Use of vitellogenin mRNA as a biomarker for endocrine disruption in feral and cultured fish. Anal Bioanal Chem. 2004;378:670-675
26. Tingaud-Sequeira A, Knoll-Gellida A, André M, Babin PJ. Vitellogenin expression in white adipose tissue in female teleost fish. BiolReprod. 2012Feb14;86(2):38
27. Cleveland BM, Weber GM. Effects of steroid treatment on growth, nutrient partitioning, and expression of genes related to growth and nutrient metabolism in adult triploid rainbow trout (Oncorhynchus mykiss). DomestAnimEndocrinol. 2016Jul;56:1-12 Epub 2016 Jan 25
28. Karpova T, Danchuk S, Kolobova E, Popov KM. Characterization of the isozymes of pyruvate dehydrogenase phosphatase: implications for the regulation of pyruvate dehydrogenase activity. BiochimBiophysActa. 2003Dec1;1652(2):126-35
29. Magnadóttir B. Innate immunity of fish (overview). Fish Shellfish Immunol. 2006Feb;20(2):137-51 Review
30. Nakao M, Mutsuro J, Obo R, Fujiki K, Nonaka M, Yano T. Molecular cloning and protein analysis of divergent forms of the complement component C3 from a bony fish, the common carp (Cyprinuscarpio): presence of variants lacking the catalytic histidine. Eur J Immunol. 2000Mar;30(3):858-66
31. Jayasinghe JD, Elvitigala DA, Whang I, Nam BH, Lee J. Molecular characterization of two immunity-related acute-phase proteins: Haptoglobin and serum amyloid A from black rockfish (Sebastes schlegeli). Fish Shellfish Immunol. 2015Aug;45(2):680-8 Epub 2015 May 16
32. Løvoll M, Kilvik T, Boshra H, Bøgwald J, Sunyer JO, Dalmo RA. Maternal transfer of complement components C3-1, C3-3, C3-4, C4, C5, C7, Bf, and Df to offspring in rainbow trout (Oncorhynchus mykiss). Immunogenetics. 2006Apr;58(2-3):168-79 Epub 2006 Mar 21
33. Brown M, Hablützel P, Friberg IM, Thomason AG, Stewart A, Pachebat JA, Jackson JA. Seasonal immunoregulation in a naturally-occurring vertebrate. BMC Genomics. 2016May18;17(1):369
34. Kocmarek AL, Ferguson MM, Danzmann RG. Differential gene expression in small and large rainbow trout derived from two seasonal spawning groups. BMC Genomics. 2014Jan22;15:57
35. Dietrich MA, Westfalewicz B, Jurecka P, Irnazarow I, Ciereszko A. Isolation, characterisation and cDNA sequencing of a new form of parvalbumin from carp semen. ReprodFertil Dev. 2014Oct;26(8):1117-28
36. Dietrich MA, Nynca J, Bilińska B, Kuba J, Kotula-Balak M, Karol H, Ciereszko A. Identification of parvalbumin-like protein as a major protein of common carp(Cyprinuscarpio L) spermatozoa which appears during final stage ofspermatogenesis. Comp BiochemPhysiol B BiochemMol Biol. 2010Oct;157(2):220-7 Epub 2010 Jul 1
37. Vigliano FA, Araujo AM, Marcaccini AJ, Marengo MV, Cattaneo E, Peirone C, Dasso LG. Effects of sex and season in haematological parameters and cellular composition of spleen and head kidney of pejerrey (Odontesthesbonariensis). Fish PhysiolBiochem. 2014Apr;40(2):417-26 Epub 2013Aug 28
38. Falahatkar B, Poursaeid S, Meknatkhah B, Khara H, Efatpanah I. Long-term effects of intraperitoneal injection of estradiol-17β on the growth and physiology of juvenile stellate sturgeon Acipenserstellatus. Fish PhysiolBiochem. 2014Apr;40(2):365-73 Epub 2013 Aug 30
39. Nagdas SK, Hamilton SL, Raychoudhury S. Identification of acrosomal matrix-specific hydrolases binding proteins of bovine cauda epididymal spermatozoa. J Androl. 2010 Mar-Apr;31(2):177-87.Epub 2009 May 28
40. Lu J, Zheng M, Zheng J, Liu J, Liu Y, Peng L, Wang P, Zhang X, Wang Q, Luan P, Mahbooband S, Sun X. Transcriptomic Analyses Reveal Novel Genes with Sexually Dimorphic Expression in Yellow Catfish (Pelteobagrusfulvidraco) Brain. Mar Biotechnol (NY). 2015Oct;17(5):613-23 Epub 2015 Aug 5
41. Zeng Q, Liu S, Yao J, Zhang Y, Yuan Z, Jiang C, Chen A, Fu Q, Su B, Dunham R, Liu Z. Transcriptome Display During Testicular Differentiation of Channel Catfish (Ictalurus punctatus) as Revealed by RNA-Seq Analysis. BiolReprod. 2016 Jun 15. pii: biolreprod.116.138818. [Epub ahead of print]
42. Barrett TJ, Munkittrick KR. Seasonal reproductive patterns and recommended sampling times for sentinel fish species used in environmental effects monitoring programs in canada. Environmental Reviews. 2010;18:115-135
43. Berninger JP, Martinović-Weigelt D, Garcia-Reyero N. et al. Using transcriptomic tools to evaluate biological effects across effluent gradients at a diverse set of study sites in minnesota, USA. Environmental Science and Technology. 2014;48(4):2404-2412
44. Bahamonde PA, McMaster ME, Servos MR, Martyniuk CJ, Munkittrick KR. Molecular pathways associated with the intersex condition in rainbow darter (etheostomacaeruleum) following exposures to municipal wastewater in the grand river basin, ON, Canada. Part B. Aquatic Toxicology. 2015;159:302-316
45. Baldigo BP, George SD, Phillips PJ, Hemming JDC, Denslow ND, Kroll KJ. Potential estrogenic effects of wastewaters on gene expression in pimephalespromelas and fish assemblages in streams of southeastern New York. Environmental Toxicology and Chemistry. 2015;34(12):2803-2815
46. Knight LA, Christenson MK, Trease AJ, Davis PH, Kolok AS. The spring runoff in Nebraska's (USA) Elkhorn River watershed and its impact on two sentinel organisms. Environ Toxicol Chem. 2013Jul;32(7):1544-51.Epub 2013 May 28
47. Ali JM, Kolok AS. On-site, serial exposure of female fathead minnows to the Elkhorn River, Nebraska, USA, spring agrichemical pulse. Environ Toxicol Chem. 2015Jun;34(6):1354-61 Epub 2015 Apr 9
48. Zhang Y, Krysl RG, Ali JM, Snow DD, Bartelt-Hunt SL, Kolok AS. Impact of Sediment on Agrichemical Fate and Bioavailability to Adult Female Fathead Minnows: A Field Study. Environ Sci Technol. 2015Aug4;49(15):9037-47 Epub 2015 Jul 21
49. Garcia-Reyero N, Adelman IR, Martinović D, Liu L, Denslow ND. Site-specific impacts on gene expression and behavior in fathead minnows (pimephalespromelas) exposed in situ to streams adjacent to sewage treatment plants. BMC Bioinformatics. 2009;10(Suppl 11):S11
50. Sellin MK, Snow DD, Akerly DL, Kolok AS. Estrogenic compounds downstream from three small cities in eastern nebraska: Occurrence and biological effect. Journal of the American Water Resources Association. 2009;45(1):14-21
51. Werner J, Ouellet JD, Cheng CS. et al. Pulp and paper mill effluents induce distinct gene expression changes linked to androgenic and estrogenic responses in the fathead minnow (pimephalespromelas). Environmental Toxicology and Chemistry. 2010;29(2):430-439
52. Ankley GT, Bencic DC, Breen MS. et al. Endocrine disrupting chemicals in fish: Developing exposure indicators and predictive models of effects based on mechanism of action. Aquatic Toxicology. 2009;92(3):168-178
53. Kramer VJ, Etterson MA, Hecker M. et al. Adverse outcome pathways and ecological risk assessment: Bridging to population-level effects. Environmental Toxicology and Chemistry. 2011;30(1):64-76
54. OECD Economic Surveys: Chile 2013, OECD Publishing, Paris. OECD. http://dx.doi.org/10.1787/eco_surveys-chl-2013-en
55. Chiang G, Munkittrick KR, McMaster ME, Barra R, Servos M. Regional cumulative effects monitoring framework: Gaps and challenges for the biobío river basin in south central chile [Marco conceptual de Monitoreo Regional de EfectosAcumulativos: Brechas y desafíos para la Cuenca del rio Biobío en el centro-sur de Chile]. Gayana. 2014;78(2):109-119
56. Chappie DJ, Burton GAJ. Applications of aquatic and sediment toxicity testing in situ. Soil Sediment Contam. 2000;9(3):219-45
57. Aranda PS, LaJoie DM, Jorcyk CL. Bleach gel: a simple agarose gel for analyzing RNA quality. Electrophoresis. 2012Jan;33(2):366-9
58. FastQC: a quality control tool for high throughput sequence data. Andrews S. http://www.bioinformatics.babraham.ac.uk/projects/fastqc
59. Schmieder R, Edwards R. Quality control and preprocessing of metagenomic datasets. Bioinformatics. 2011Mar15;27(6):863-4 Epub 2011 Jan 28
60. Grabherr MG, Haas BJ, Yassour M, Levin JZ, Thompson DA, Amit I, Adiconis X, Fan L, Raychowdhury R, Zeng Q, Chen Z, Mauceli E, Hacohen N, Gnirke A, Rhind N, di Palma F, Birren BW, Nusbaum C, Lindblad-Toh K, Friedman N, Regev A. Full-length transcriptome assembly from RNA-seq data without a reference genome. Nat Biotechnol. 2011May15;29(7):644-52
61. Haas BJ, Papanicolaou A, Yassour M, Grabherr M, Blood PD, Bowden J, Couger MB, Eccles D, Li B, Lieber M, Macmanes MD, Ott M, Orvis J, Pochet N, Strozzi F, Weeks N, Westerman R, William T, Dewey CN, Henschel R, Leduc RD, Friedman N, Regev A. De novo transcript sequence reconstruction from RNA-seq using the Trinity platform for reference generation and analysis. Nat Protoc. 2013Aug;8(8):1494-512 Epub 2013 Jul 11
62. Henschel R, Lieber M, Wu L, Nista PM, Haas BJ, LeDuc R. Trinity RNA-Seq assembler performance optimization. XSEDE. 2012 Proceedings of the 1st Conference of the Extreme Science and Engineering Discovery Environment: Bridging from the eXtreme to the campus and beyond. ISBN: 978-1-4503-1602-6 doi: 10.1145/2335755.2335842
63. Camacho C, Coulouris G, Avagyan V, Ma N, Papadopoulos J, Bealer K, Madden TL. BLAST+: architecture and applications. BMC Bioinformatics. 2009Dec15;10:421
64. Castresana J. Selection of conserved blocks from multiple alignments for their use in phylogenetic analysis. Molecular biology and evolution. 2000;17(4):540-552
65. Talavera G, Castresana J. Improvement of phylogenies after removing divergent and ambiguously aligned blocks from protein sequence alignments. Systematic biology. 2007;56(4):564-577
66. Thompson JD, Higgins DG, Gibson TJ. CLUSTAL W: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res. 1994Nov11;22(22):4673-80
67. Conesa A, Götz S, García-Gómez JM, Terol J, Talón M, Robles M. Blast2GO: a universal tool for annotation, visualization and analysis in functional genomics research. Bioinformatics. 2005Sep15;21(18):3674-6 Epub 2005 Aug 4
68. Langmead B, Trapnell C, Pop M, Salzberg SL. Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol. 2009;10(3):R25. Epub 2009 Mar 4
69. Li B, Dewey CN. RSEM: accurate transcript quantification from RNA-Seq data with or without a reference genome. BMC Bioinformatics. 2011Aug4;12:323
70. FishBase; World Wide Web electronic publication. Froese R and Pauly D. www.fishbase.org
Author contact
Corresponding author: Dr. Paul H. Davis. pdavisedu

 Global reach, higher impact
Global reach, higher impact