ISSN: 1839-9940
J Genomics 2018; 6:34-40. doi:10.7150/jgen.22460 This volume Cite
Research Paper
In-silico Designing and Testing of Primers for Sanger Genome Sequencing of Dengue Virus Types of Asian Origin
1. Desert Medicine Research Centre, Indian Council of Medical research, Jodhpur, India-342005
2. Present Address: Amity Institute of Virology & Immunology (AIVI), Amity University, Noida, U.P., India- 201313
3. Present Address: National Institute of Malaria Research, Secotr-8, Dwarika, New Delhi -110077
4. All India Institute of Medical Sciences, Jodhpur, India
5. Department of Neuroscience, Dr. SN Medical College, Jodhpur, India-342001
6. Department of Medicine, Dr. SN Medical College, Jodhpur, India-342001
Abstract
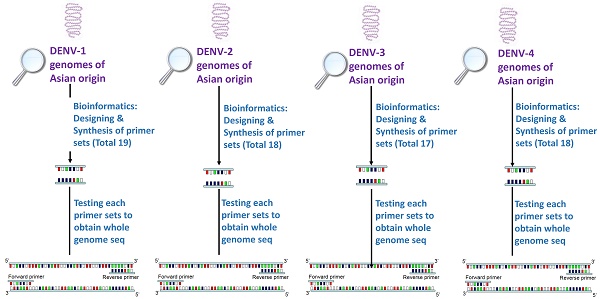
Rarity in reporting whole genome sequence of Dengue virus from dengue endemic countries leaves lacunae in understanding regional pattern of virus mutation and ultimately leading to non-understanding of transmission pattern and clinical outcomes emerging at regional levels. Due to inter-serotype genomic similarity and intra-serotype genomic diversity, appropriate designing of primer pairs appears as an exhaustive exercise. Present paper reports new Dengue virus type-specific primer which may help in characterizing virus specific to Asian origin. Genomes of dengue virus serotypes of Asian region were searched and using advanced bioinformatics tools, serotype specific primers were designed and tested for their targeted amplification efficiency. 19 primers sets for DENV-1, 18 primer sets for DENV-2, 17 for DENV-3 and 18 for DENV-4 were designed. In-silico and experimental testing of the designed primers were performed on virus isolated from both clinical isolates and passaged cultures. While all 17 and 18 primer sets of DENV-3 and DENV-2 respectively yielded good quality sequencing results; in case of DENV-4, 16 out of 18 primer sets and in DENV-1, 16 out of 19 primer sets yielded good results. Average sequencing read length was 382 bases and around 82% nucleotide bases were Phred quality QV20 bases (representing an accuracy of circa one miscall every 100 bases) or higher. Results also highlighted importance of use of primer development algorithm and identified genomic regions which are conservative, yet specific for developing primers to achieve efficiency and specificity during experiments.
Keywords: Flavivirus, Mutation, Oligonucleotide, BLAST algorithm, Genetic variation

 Global reach, higher impact
Global reach, higher impact