ISSN: 1839-9940
J Genomics 2019; 7:14-17. doi:10.7150/jgen.31170 This volume Cite
Short Research Communication
Genome Sequencing and Annotation of Bacillus subtilis UBBS-14 to Ensure Probiotic Safety
Center for Research and Development, Unique Biotech Limited, Hyderabad - 500 078, India
Received 2018-11-2; Accepted 2019-1-8; Published 2019-1-29
Abstract
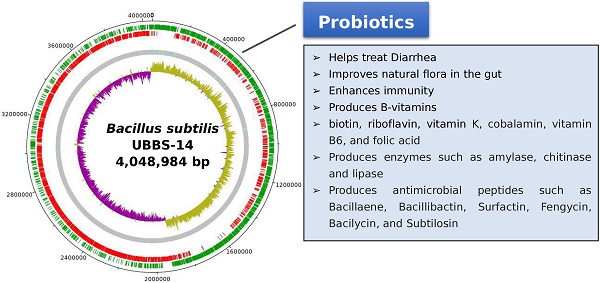
Bacillus subtilis is a rod shaped, gram positive, spore producing bacterium. They are the normal flora of gastrointestinal tract of humans and it is the best characterized model organism for endospore formation. It has the ability to withstand environmental stress, and synthesize beneficial compounds, therefore, it is recognized as a high-quality probiotic supplement. To ensure the probiotic safety and the efficiency, we report the whole genome sequence (WGS) of Bacillus subtilis UBBS-14 strain. The draft genome sequence of Bacillus subtilis UBBS-14 consists of 4,048,984 bp and 4,017 genes, respectively. Bacillus subtilis UBBS-14 does not carry any antibiotic resistant genes containing plasmid, nor it contains any harmful putative virulence factors coding genes, therefore, it confirms the probiotic safety of the respective strain at genome level.
Introduction
Bacillus subtilis are the most widespread bacteria in soil, water, air and gastrointestinal tract of mammals. They are Gram-positive, rod shaped bacteria, known for their ability to produce a robust spore [1-5]. The complete genome sequence of Bacillus subtilis has deduced the operating mechanisms of the organism [6]. It is well known for its ability to produce secondary metabolites, bacteriocins, enzymes, antibiotics and vaccines [1,7-12]. Here, we make an announcement of draft genome of Bacillus subtilis UBBS-14 strain, which was isolated from fermented food at Unique Biotech Limited, Hyderabad, India.
Materials and Methods
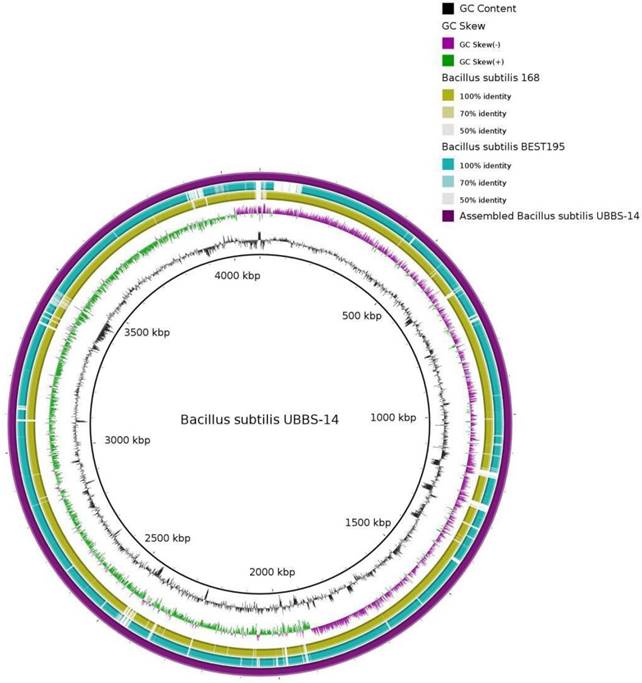
Bacillus subtilis UBBS-14 was isolated from the fermented food, the isolate was subjected for DNA isolation and 16S rDNA identification using universal forward 27F (5' AGAGTTTGATCMTGGCTCAG 3') and reverse 1429R (5'TACGGYTACCTTGTTACGACTT3') primers. Following the species identification, the strain was outsourced for WGS (Whole Genome Sequencing) to Genotypic Technology Pvt. Ltd., Bengaluru, India. The whole-genome sequencing was performed using the Illumina MiSeq platform, with a paired-end library; A total of 2,554,530 paired-end raw reads of 150-bp length on average (genome coverage of 189x) were sequenced, out of which, 2,193,966 high quality paired-end reads, with 80% read length scoring Phred quality score of 30 and above, were assembled into 34 Scaffolds by employing de novo genome assembler SPAdesv3.11.1 [13] and scaffolder SSPACE-STANDARD-v.-3.0. [14]. A draft genome map was drawn by using DNA plotter [15], the origin and terminus were plotted through GenSkew server, and the statistics of genome assembly was calculated by NGS QC Toolkit [16]. The genome sequence was annotated by RAST server [17] and by the NCBI's Prokaryotic Genomes Annotation Pipeline (PGAP) [18]. The gene prediction was carried out using Prodigal program and the predicted proteins were searched for similarity against Uniprot protein database using Blastp program [19], following pathway identification by KEGG-KAAS server. The genome was screened to determine the putative virulence factors (VFDB database) [20], plasmid (PlasmidFinder 2.0) [21], and antibiotic resistant genes (ARDB) [22]. BRIG (Blast Ring Image Generator) plot was constructed to compare the assembled genome of Bacillus subtillis UBBS-14 with the standard reference strains Bacillus subtilis subsp., subtilis 168 and Bacillus subtilis BEST195 [23].
Results and Discussion
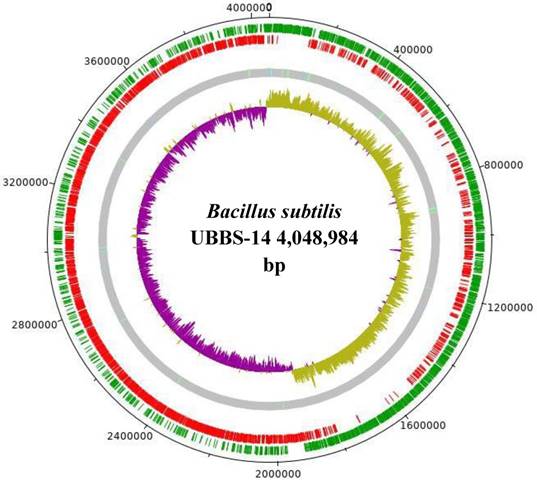
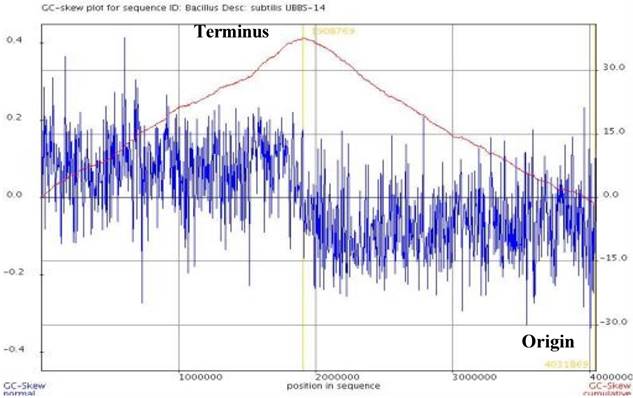
The draft genome consists of 4,048,984 bp, with the largest assembled scaffold of 5,60,413 bp length, GC % content of 43.69 (Figure: 1), the origin of replication is located at 19,08,765 bp, whereas, the terminus is located at 40,31,869 bp (Figure: 2). A total of 4,017 genes were predicted, of which 3,845 are coding sequences (CDS), 7 are rRNAs, and 96 are tRNAs. The strain is predicted to encode for about 393 proteins involved in carbohydrate metabolism and 277 proteins involved in amino acid metabolism, and 44 putative proteins for xenobiotics biodegradation and metabolism.
The genes encoding for putative virulence factors such as hemolysin BL, non hemolytic enterotoxin NHE, enterotoxin T, cytotoxin T and cereulide were not found. The genome has no plasmid, the antibiotic resistant genes against Beta-lactam, Fluoroquinolone, Fosfomycin, Fusidic acid, MLS-Macrolide-Lincosamide-Streptogramin B, Nitroimidazole, Phenicol, Rifampicin, Sulphonamide, Tetracycline, Trimethoprim, Glycopeptide were not found. The genome has Aminoglycoside 6-adenylyltransferase coding gene, the length and amino acid sequence of this particular enzyme is 100 percent identical with the the product of 'aadk' gene of Bacillus subtilis subsp., subtilis 168 strain, which has reported to confer low-level resistance to streptomycin [24].
The Bacillus subtilis UBBS-14 strain contains genes involved in the biosynthesis of biotin, riboflavin, vitamin K, cobalamin, vitamin B6, folic acid, which leads it to be an nutrition probiotic. It also codes for the antimicrobial peptides such as Bacillaene, Bacillibactin, Surfactin, Fengycin, Bacilycin, and Subtilosin A. The assembled genome of Bacillus subtilis UBBS-14 strain showed 99% sequence identity and 90% coverage with reference Bacillus subtilis Best 195, and 99% sequence identity and 94% coverage with the reference Bacillus subtilis subsp., subtilis 168 (Figure 3). The genome analysis of Bacillus subtilis UBBS-14 shows that it does not contain any virulence and antibiotic resistant genes containing plasmid, which ensure its probiotic safety, further studies on the phenotypic characterization of the strain shall give better insights to understand the findings of its genomic characteristics.
Draft genome of Bacillus subtilis UBBS-14. Green circle - CDS on the plus strand, red circle - CDS on the minus strand, grey circle - Scaffolds with light green bars of tRNA genes, light blue bars of repeat regions, inner circle is GC-skew plot (magenta portion - GC-skew positive, olive green portion - GC-skew negative).
GC-Skew plot for Bacillus subtilis UBBS-14
BRIG based on homology with Reference Strains
Accession Numbers
This whole-genome shotgun project of Bacillus subtilis UBBS-14 has been deposited in DDBJ/EMBL/GenBank under the accession number RDEZ00000000. The version described in this paper is the first version, RDEZ01000000.
Acknowledgements
The authors acknowledge the “Genotypic Technology Pvt. Ltd”, Bengaluru, India for their services.
Competing Interests
The authors have declared that no competing interest exists.
References
1. Sonnenschein AL, Losick R, Hoch JA. Bacillus subtilis and d other gram-positive bacteria: biochemistry, physiology, and molecular genetics. Washington DC, USA: American Society for Microbiology. 1993
2. Garbeva P, van Veen JA, van Elsas JD. Predominant Bacillus spp. In agricultural soil under different management regimes detected via PCR-DGGE. Microb Ecol. 2003;45:302-316
3. Nicholson W. Roles of Bacillus endospores in the environment. Cell Mol Life Sci. 2002;59:410-416
4. Ivanova EP, Vysotskii MV, Svetashev VI, Nedashkovskaya OI, Gorshkova NM. et al. Characterization of Bacillus strains of marine origin. Int Microbiol. 1999;2:267-271
5. Andersson AM, Weiss N, Rainey F, Salkinoja-Salonen MS. Dust-borne bacteria in animal sheds, schools and children's day care centres. J Appl Microbiol. 1999;86:622-634
6. Kunst F, Ogasawara N, Moszer I, Albertini AM, Alloni G. et al. The complete genome sequence of the gram-positive bacterium Bacillus subtilis. Nature. 1997;390:249-256
7. Berdy J. Bioactive microbial metabolites. J Antibiot. 2005;58:1-26
8. Stein T. Bacillus subtilis antibiotics: structures, syntheses and specific functions. Mol Microbiol. 2005;56:845-857
9. Ferrari E, Jarnagin AS, Schmidt BF. Commercial production of extra cellular enzymes. Bacillus subtilis and Other Gram-Positive Bacteria. Sonenshein AL Hoch JA, Losick R, eds. Washington DC, USA: American Society for Microbiology. 1993:917-937
10. Olmos-Soto J, Contreras-Flores R. Genetic system constructed to overproduce and secrete proinsulin in Bacillus subtilis. Appl Microbiol Biotechnol. 2003;62:369-373
11. Westers L, Westers H, Quax WJ. Bacillus subtilis as cell factory for pharmaceutical proteins: a biotechnological approach to optimize the host organism. Biochim Biophys Acta. 2004;1694:299-310
12. Valdez A, Yepiz-Plascencia G, Ricca E, Olmos J. First Litopenaeus vannamei WSSV 100% oral vaccination protection using CotC::Vp26 fusion protein displayed on Bacillus subtilis spores surface. J Appl Microbiol. 2014;117:347-357
13. Anton Bankevich, Sergey Nurk, Dmitry Antipov, Alexey A Gurevich, Mikhail Dvorkin, Alexander S. Kulikov, Valery M Lesin, Sergey I Nikolenko, Son Pham, Andrey D Prjibelski, Alexey V Pyshkin, Alexander V Sirotkin, Nikolay Vyahhi, Glenn Tesler, Max A Alekseyev, and Pavel A Pevzner, SPAdes: A New Genome Assembly Algorithm and Its Applications to Single-Cell Sequencing. J Comput Biol. 2012;19(5):455-477
14. Martin Hunt, Chris Newbold, Matthew Berriman, Thomas D Otto. A comprehensive evaluation of assembly scaffolding tools. Genome Biol. 2014;15(3):R42
15. Carver T, Thomson N, Bleasby A. et al. DNAPlotter: circular and linear interactive genome visualization. Bioinformatics. 2008;25:119-120
16. Ravi K Patel, Mukesh Jain. NGS QC Toolkit: A Toolkit for Quality Control of Next Generation Sequencing Data. PLoS One. 2012;7(2):e30619
17. Aziz RK, Bartels D, Best AA. et al. The RAST Server: rapid annotations using subsystems technology. BMC Genomics. 2008;9:75
18. Tatusova T, DiCuccio M, Badretdin A. et al. NCBI prokaryotic genome annotation pipeline. Nucleic Acids Res. 2016;44:6614-6624
19. Hyatt Doug. et al. Prodigal: prokaryotic gene recognition and translation initiation site identification. BMC bioinformatics. 2010;11:119
20. Chen LH, Zheng DD, Liu B, Yang J, Jin Q. VFDB 2016: hierarchical and refined dataset for big data analysis-10 years on. Nucleic Acids Res. 2016;44:D694-697
21. Carattoli A, Zankari E, García-Fernández A, Voldby Larsen M, Lund O, Villa L, Møller Aarestrup F, Hasman H. In silico detection and typing of plasmids using PlasmidFinder and plasmid multilocus sequence typing. Antimicrob Agents Chemother. 2014;58(7):3895-903
22. Liu Bo, Mihai Pop. ARDB—antibiotic resistance genes database. Nucleic Acids Res. 2009;37(Database issue):D443-7
23. Nabil-Fareed Alikhan, Nicola K Petty, Nouri L Ben Zakour, Scott A Beatson. BLAST Ring Image Generator (BRIG): simple prokaryote genome comparisons. BMC Genomics. 2011;12:402
24. Noguchi N, Sasatsu M, Kono M. Genetic mapping in Bacillus subtilis 168 of the aadK gene which encodes aminoglycoside 6-adenylyltransferase. FEMS Microbiol Lett. 1993;114(1):47-52
Author contact
Corresponding author: Dr. Ayesha Sulthana. Email: ayeshasulthanaacom

 Global reach, higher impact
Global reach, higher impact