ISSN: 1839-9940
J Genomics 2020; 8:11-15. doi:10.7150/jgen.38461 This volume Cite
Research Paper
Draft Genome Sequence of the Symbiotic Frankia sp. strain B2 isolated from root nodules of Casuarina cunninghamiana found in Algeria
1. Laboratoire de Biologie du Sol, Faculté des Sciences Biologiques, Université des Sciences et de la Technologies Houari Boumediene (USTHB), BP32 El Alia - Bab Ezzouar Algiers, Algeria.
2. University of New Hampshire, 46 College Rd., Durham, New Hampshire, USA, 03824-2617
3. Laboratoire des Symbioses Tropicales et Méditerranéennes (IRD/INRA/CIRAD/Université de Montpellier/Supagro), 34398 Montpellier Cedex 5, France.
4. Laboratoire Molécules de Communication et Adaptation des Microorganismes (MCAM) UMR 7245 CNRS-MNHN), Museum national d'Histoire naturelle, Centre National de la Recherche Scientifique (CNRS), CP 54, 57 rue Cuvier, 75005 Paris, France
Received 2019-7-16; Accepted 2019-11-22; Published 2020-1-19
Abstract

Frankia sp. strain B2 was isolated from Casuarina cunninghamiana nodules. Here, we report the 5.3-Mbp draft genome sequence of Frankia sp. strain B2 with a G+C content of 70.1 % and 4,663 candidate protein-encoding genes. Analysis of the genome revealed the presence of high numbers of secondary metabolic biosynthetic gene clusters.
Keywords: actinorhizal symbiosis, host-microbe interactions, nitrogen fixation, Casuarinaceae, Frankia, land reclamation, genomes
Introduction
Actinobacteria of the genus Frankia are Gram positive filamentous bacteria that are able to fix molecular nitrogen in free living state or in symbiosis with their host plant [1, 2]. These bacteria establish a nitrogen-fixing symbiosis with a diverse variety of plant species, collectively named actinorhizal plants, which include 8 dicotyledonous plant families, 24 genera and over 220 species. The mutualistic association is referred to as the actinorhizal symbiosis and results in the formation of a root nodule structure. The bacteria are housed within plant cells in the nodule which allows for the trophic exchange between the two partners. The bacteria reduce atmospheric nitrogen to ammonia that is supplied to the host plant, which in return provides carbon compounds from photosynthesis to the bacteria. Because of the symbiosis, actinorhizal plants can colonize poor and degraded soils and thrive in inhospitable and harsh habitats [2]. Actinorhizal plants are pioneer species that allow the succession of other plant communities by providing organic matter, a fundamental matrix for the dynamics and biodiversity of terrestrial ecosystems. There is currently a renewing interest for actinorhizal symbiosis due to its significant contribution to global soil amendment in combined nitrogen (more than 15%) [3].
Based on the recent molecular phylogenetic studies, Frankia strains are classified into four major clusters [4-6] that reflect host plant range. Cluster 1 consists of Frankia strains that associate with host plants in the Casuarinaceae, Betulaceae and Myricaceae families, while members of cluster 2 are infective on Rosaceae, Coriariaceae, Datiscaceae, and the genus Ceanothus (Rhamnaceae). Cluster 3 are the most promiscuous and are infective on Elaeagnaceae, Rhamnaceae, Myricaceae, Gynmnostoma, and occasionally the genus Alnus. Cluster 4 consists of “atypical” Frankia strains that are unable to re-infect actinorhizal host plants or form ineffective nonnitrogen-fixing root nodule structures. Cluster 1 is further divided into subclades. Subclade Ic includes strains limited to Casuarina and Allocasuarina and Myricaceae host plants.
Actinorhizal species include Casuarina spp., tropical trees native in Australia, Southeast Asia and Oceania [7]. These woody plants are well adapted to drought, heat, salinity, polluted soils and can withstand multiple varieties of environments [2]. This property is one reason why they have been massively planted in several regions of the globe for land reclamation, prevention of erosion, crop protection and fighting against desertification, tsunamis and typhoons [7]. In Algeria, like in all the Maghreb, Casuarina trees were introduced in the 19th century and are currently found widespread in all bioclimatic zones of the country ranging from the coastal zone to the Saharan areas. Today, the propagation of Casuarina trees occurs mostly from plantlets produced in nurseries via seeds or by cutting. As a part of a project that aims to reassess the identity, the distribution and the relative abundance of Casuarina trees in Algeria, we were interested in investigating the prevalence of actinorhizal symbiosis in nurseries from different regions of the country, and to examine whether the symbiotic status can help the installation of the plantlets in natural environments. For this purpose, we have collected nodules samples from young Casuarina trees from Algerian nurseries and the symbiotic Frankia strain was isolated.
Isolation of Frankia strain B2
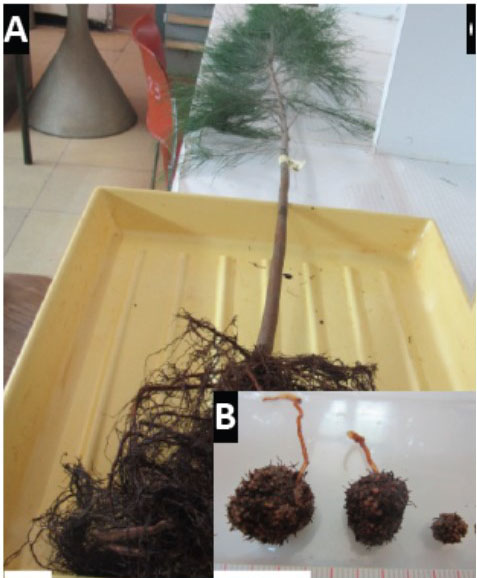
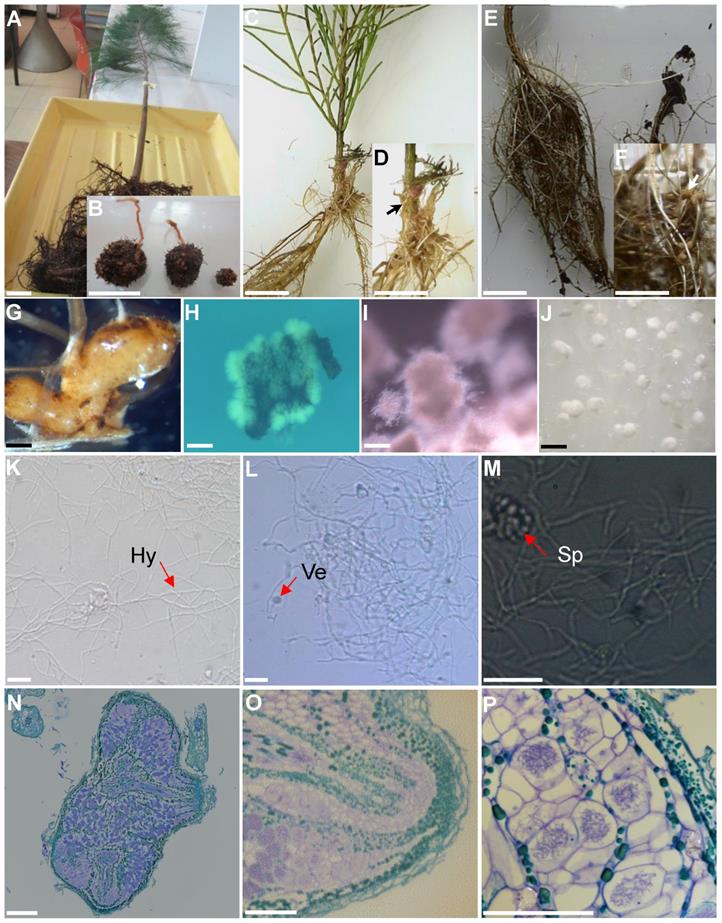
Frankia strain B2 was isolated in two-step process from nodules collected from Casuarina cunninghamiana seedlings growing in a nursery located at Souk El Tenine (District of Bejaia, Algeria). For the first step, the collected nodules were crushed and used as an inoculum on Casuarina glauca plants growing hydroponically in N-free BD medium [8] in a culture chamber under controlled conditions (25° C, 75% of relative air humidity and 16 h of photoperiod. After 8 weeks, root nodules were observed and harvested. For the second step, harvested nodules were washed, fragmented and surface-sterilized by immersion in a 30% H202 solution for 30 min based on protocol described previously [9]. Sterilized nodule fragments were inoculated onto the surface of different solid growth media including BAP [10], DPM (Defined Propionate Minimal Medium) [11] or modified QMOD [12] under nitrogen-free conditions (without yeast extract and peptone for QMOD). Plates were incubated in dark at 28°C. After 4-6 weeks, Frankia hyphae developed around the nodule fragments inoculated on BAP medium and these colonies were transferred into liquid BAP growth medium. Figure 1 shows the different stages of the isolation process and photomicrographs show typical Frankia features. Frankia has three different morphogenetic forms; vegetative hyphae (Hy), vesicles (Ve), the site of nitrogen fixation and sporangia containing spores (Sp). All three types of cell structures were produced by Frankia strain B2 (Fig. 1K-M). Frankia strain B2 was able to re-infect C. cunninghamiana and the nodules produced (Fig. 1N-O) showed a higher level of nitrogenase activity compared to C. cunninghamiana nodules with Frankia casuarinae strain CcI3, the type strain [13] (Fig. 2). The acetylene reduction activity (ARA) was used to determine nitrogenase activity of C. cunninghamiana [14]. Because Frankia strain B2 had these traits and it represented an Algerian isolate, we chose to sequence its genome.
Isolation of Frankia strain B2. (A) A nodulated young Casuarina cunninghamiana grown in a nursery of Souk El Tenine (District of Bejaia, Algeria). (B) C. cunninghamiana root nodules. (C) Nodulation of C. cunninghamiana after inoculation with crushed nodules previously harvested from C. glauca growing hydroponically in N-free BD medium (8) for 8 weeks in growth chamber at 25° C with 75% of relative air humidity and 16 h of photoperiod. (D) Close up of C, showing nodule (arrow). (E) C. glauca nodulation after inoculation with crushed nodules harvested from C. cunninghamiana grown in the nursery. (F) Close up of panel E, showing nodule (arrow). (G) Casuarina glauca young nodule used for Frankia B2 isolation. (H) Development of Frankia B2 from a surface-sterilized C. glauca nodule fragment cultivated on BAP solid medium. (I) Cultures of Frankia B2 grown in BAP liquid medium. (J) Frankia B2 colonies cultivated on BAP solid medium. (K) Frankia B2 isolated from colonies (see panel J) and cultivated in BAP liquid medium. Hyphae are observed (Hy). (L, M) Frankia B2 liquid culture showing vesicles (Ve) and sporangia (Sp). (N) Longitudinal section of a C. cunninghamiana nodule stained with toluidine blue. (O) Close up of panel N showing a nodule lobe. (P) Close up of panel O showing cortical infected cells. Scale bars: A-F = 20 mm; G-J = 5 mm; K-M = 100 µm; N-O = 100 µm.
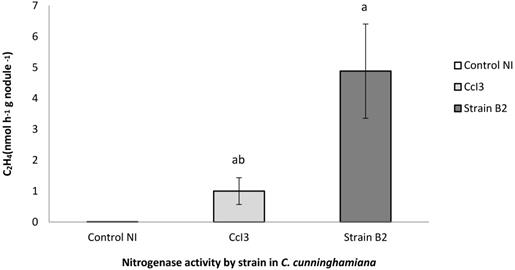
Nitrogenase activity of nodules induced by Frankia strain B2. The acetylene reduction activity (ARA) was used to determine nitrogenase activity of C. cunninghamiana [14]. Nodules of C. cunninghamiana were induced with Frankia strain B2 or F. casuarinae strain CcI3. The uninoculated C. cunninghamiana plants (NI) were included as a control. Values represent the mean of several measurements (Control NI, N = 4; F. casuarinae strain CcI3, N = 4; and Frankia strain B2, N = 23), Error bars indicate standard error of the mean ANOVA-LSD analysis shows that a significant difference among the samples.
Sequencing of Frankia strain B2
Sequencing of the draft genome of Frankia sp. strain B2 was performed at the Hubbard Center for Genome Studies (University of New Hampshire, Durham, NH) using Illumina technology techniques [15]. High quality gDNA of Frankia sp. strain B2 was extracted using CTAB method [16]. A standard Illumina shotgun library was constructed and sequenced using the Illumina HiSeq2500 platform, which generated 4,247,110 reads (260-bp insert size) totaling 965 Mbp. The Illumina sequence data were trimmed by Trimmomatic version 0.36 [17], and assembled using Spades version 3.10 [18]. The final draft assembly for Frankia sp. strain B2 consisted of 145 contigs with an N50 contig size of 103.6 kb and 176X coverage of the genome. The final assembled genome contained a total sequence length of 5,331,433 bp with a G+C content of 70.12%.
The assembled Frankia sp. strain B2 genome was annotated via the NCBI Prokaryotic Genome Annotation Pipeline (PGAP), and resulted in 4,663 candidate protein-encoding genes, 41 tRNA and 5 rRNA. The genome features of Frankia sp. strain B2 are similar to other cluster 1c genomes (Table 1) including F. casuarinae strain CcI3T [13]. Phylogenetic analysis of the 16S rDNA shows that Frankia sp. strain B2 groups with the cluster 1c strains (Figure S1) and further confirmed by dendrogram of the entire genomes (Figure S2). The genome also contained a nif, 2 hup, and 1 shc operons encoding the nitrogenase, hydrogenase uptake enzymes, and the hopanoid biosynthetic pathway, respectively. The operons were organized similar to those reported for Frankia cluster 1c genomes [19]. The pan-genome of Frankia cluster 1c consisted of 4,736 genes including a core genome of 3,107 genes. Figure S3 shows a Venn diagram of the orthologs shared among six Frankia cluster 1c strains.
Genome features of Frankia sp. strain B2 and other Frankia strains isolated from Casuarina root nodules.
| Strain | Source | Location1 | Size (Mb) | No. of Contig(s) | G+C (%) | No. of CDS | No. of rRNA | No. of tRNA |
|---|---|---|---|---|---|---|---|---|
| B2 | This study | Algeria | 5.33 | 145 | 70.1 | 4,663 | 5 | 45 |
| KB5 | [22] | Australia | 5.46 | 420 | 70.0 | 4,958 | 6 | 45 |
| CcI3 | [23] | USA | 5.43 | 1 | 70.1 | 4,598 | 6 | 46 |
| CeD | [24] | Senegal | 5.00 | 120 | 70.1 | 4,403 | 7 | 45 |
| Allo2 | [25] | Uruguay | 5.33 | 110 | 69.8 | 4,838 | 7 | 46 |
| Thr | [26] | Egypt | 5.31 | 171 | 70.0 | 4,805 | 5 | 46 |
| BMG5.23 | [27] | Tunisia | 5.27 | 167 | 70.0 | 4,747 | 9 | 47 |
| CcI6 | [28] | Egypt | 5.39 | 138 | 67.6 | 4,902 | 9 | 46 |
| BR | [29] | Brazil | 5.23 | 180 | 70.0 | 4,777 | 5 | 46 |
1 The source of the isolate.
Bioinformatic analysis of this genome by the use of the AntiSMASH program [20] revealed the presence of high numbers of secondary metabolic biosynthetic gene clusters, which is consistent with previous results for other Frankia genomes including subcluster Ic [19, 21]. Table 2 shows a comparison of the various profiles of different Casuarina isolates for these secondary metabolic biosynthetic gene clusters. Although the majority of these secondary metabolic biosynthetic gene clusters were shared among the F. casuarinae genomes, the Frankia sp. strain B2 genome contained five unique nonribosomal peptide synthase (NRPS) clusters that were completely novel without homologues to other microbes but had minimal information on the chemical structures of the natural products. Predicted monomers for some of these unique NRPS clusters were identified, but no structure could be predicted from this algorithm.
Biosynthetic gene clusters for natural products found in the genomes from Casuarina Frankia strains.
| Strain | No. of Biosynthetic gene clusters 1 | NRPS 2 | PKS 3 | Terpene | Siderophore | Bacteriocin | Lantipeptide |
|---|---|---|---|---|---|---|---|
| B2 | 31 | 6 | 9 | 4 | 1 | 0 | 6 |
| KB5 | 34 | 4 | 9 | 6 | 1 | 1 | 4 |
| CcI3 | 29 | 3 | 5 | 4 | 1 | 3 | 6 |
| CeD | 30 | 7 | 7 | 4 | 1 | 1 | 4 |
| Allo2 | 32 | 7 | 9 | 4 | 1 | 3 | 5 |
| Thr | 33 | 6 | 7 | 4 | 1 | 1 | 6 |
| BMG5.23 | 31 | 8 | 6 | 4 | 1 | 2 | 4 |
| CcI6 | 33 | 8 | 8 | 4 | 1 | 3 | 5 |
| BR | 29 | 5 | 5 | 4 | 1 | 2 | 5 |
In summary, the Frankia sp. strain B2 genome has revealed an interesting potential for secondary metabolites pathways and natural product profile and serves as another representative of Frankia cluster 1c.
Nucleotide sequence accession numbers
This whole-genome shotgun sequence has been deposited at DDBJ/EMBL/GenBank under the accession number SOPN00000000.1. The version described in this paper is the first version, SOPN01000000.
Supplementary Material
Supplementary figures.
Acknowledgements
We thank J. Fardoux for help with ARA measurements. This study was funded by the Algerian Ministry of Higher Education and Scientific Research (MESRS), the French National Research Institute for Sustainable Development (IRD), the New Hampshire Agricultural Experimental Station, the Agriculture and Food Research Initiative Grant 2015-67014-22849 from the USDA National Institute of Food and Agriculture Hatch 022821 (LST), the College of Life Science and Agriculture at the University of New Hampshire-Durham. Sequencing was performed on an Illumina HiSeq2500 purchased with an NSF MRI Grant: DBI-1229361 to WK Thomas. This is Scientific Contribution Number 2822 from the New Hampshire Agricultural Experimental Station.
Competing Interests
The authors have declared that no competing interest exists.
References
1. Normand P, Benson DR, Berry AM, Tisa LS. Family Frankiaceae. Rosenberg E, DeLong EF, Lory S, Stackebrandt E, Thompson F, editors. The Prokaryote - Actinobacteria Berlin Heidelberg: Springer-Verlag. 2014 p. 339-56
2. Ngom M, Oshone R, Diagne N, Cissoko M, Svistoonoff S, Tisa LS. et al. Tolerance to environmental stress by the nitrogen-fixing actinobacterium Frankia and its role in actinorhizal plants adaptation. Symbiosis. 2016;70:17-29
3. Dommergues YR. Contribution of actinorhizal plants to tropical soil productivity and rehabilitation. Soil Biol Biochem. 1997;29:931-41
4. Normand P, Orso S, Cournoyer B, Jeannin P, Chapelon C, Dawson J. et al. Molecular phylogeny of the genus Frankia and related genera and emendation of the family Frankiaceae. Int J Syst Bacteriol. 1996;46:1-9
5. Ghodhbane-Gtari F, Nouioui I, Chair M, Boudabous A, Gtari M. 16S-23S rRNA Intergenic Spacer Region Variability in the Genus Frankia. Microb Ecol. 2010;60:487-95
6. Nouioui I, Ghodhbane-Gtari F, Beauchemin NJ, Tisa LS, Gtari M. Phylogeny of members of the Frankia genus based on gyrB, nifH and glnII sequences. Anton Leeuw Int J G. 2011;100:579-87
7. Council NR. Casuarina: nitrogen-fixing trees for adverse sites. Washington, DC, USA: National Academic Press. 1984
8. Svistoonoff S, Sy MO, Diagne N, Barker DG, Bogusz D, Franche C. Infection-Specific Activation of the Medicago truncatula Enod11 Early Nodulin Gene Promoter During Actinorhizal Root Nodulation. Mol Plant Microbe In. 2010;23:740-7
9. Rosbrook PA, Reddell P. Isolation of Frankia from Root-Nodules of 3 Species of Casuarina. Soil Biol Biochem. 1995;27:427-9
10. Murry MA, Fontaine MS, Torrey JG. Growth Kinetics and Nitrogenase Induction in Frankia Sp ArI3 Grown in Batch Culture. Plant Soil. 1984;78:61-78
11. Baker D, Okeefe D. A Modified Sucrose Fractionation Procedure for the Isolation of Frankiae from Actinorhizal Root-Nodules and Soil Samples. Plant Soil. 1984;78:23-8
12. Lalonde M, Calvert HE. Production of Frankia hyphae and spores as an infective inoculant for Alnus species. In: (ed.) Gordon JC, Wheeler CT, Perry DA. Symbiotic Nitrogen Fixation in the Management of Temperate Forets. Corvallis, OR: Oregon State University. 1979 p. 95-110
13. Nouioui I, Ghodhbane-Gtari F, Montero-Calasanz MD, Goker M, Meier-Kolthoff JP, Schumann P. et al. Proposal of a type strain for Frankia alni (Woronin 1866) Von Tubeuf 1895, emended description of Frankia alni, and recognition of Frankia casuarinae sp. nov. and Frankia elaeagni sp. nov. Int J Syst Evol Microbiol. 2016;66:5201-10
14. Hardy RWF, Holsten RD, Jackson EK, Burns RC. Acetylene-Ethylene Assay for N2 Fixation - Laboratory and Field Evaluation. Plant Physiol. 1968;43:1185 -&
15. Bennett S. Solexa Ltd. Pharmacogenomics. 2004;5:433-8
16. Murray MG, Thompson WF. Rapid Isolation of High Molecular-Weight Plant DNA. Nucleic Acids Res. 1980;8:4321-5
17. Bolger AM, Lohse M, Usadel B. Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics. 2014;30:2114-20
18. Nurk S, Bankevich A, Antipov D, Gurevich AA, Korobeynikov A, Lapidus A. et al. Assembling single-cell genomes and mini-metagenomes from chimeric MDA products. Journal of computational biology: a journal of computational molecular cell biology. 2013;20:714-37
19. Tisa LS, Oshone R, Sarkar I, Ktari A, Sen A, Gtari M. Genomic approaches toward understanding the actinorhizal symbiosis: an update on the status of the Frankia genomes. Symbiosis. 2016;70:5-16
20. Blin K, Wolf T, Chevrette MG, Lu XW, Schwalen CJ, Kautsar SA. et al. antiSMASH 4.0-improvements in chemistry prediction and gene cluster boundary identification. Nucleic Acids Res. 2017;45:W36-W41
21. Udwary DW, Gontang EA, Jones AC, Jones CS, Schultz AW, Winter JM. et al. Significant Natural Product Biosynthetic Potential of Actinorhizal Symbionts of the Genus Frankia, as Revealed by Comparative Genomic and Proteomic Analyses. Appl Environ Microb. 2011;77:3617-25
22. Pesce C, Swanson E, Simpson S, Morris K, Thomas WK, Tisa LS. et al. Draft Genome Sequence for Frankia sp. Strain KB5 Isolated from the Root Nodules of Casuarina equisetifolia. J Genomics. 2017;5:64-7
23. Normand P, Lapierre P, Tisa LS, Gogarten JP, Alloisio N, Bagnarol E. et al. Genome characteristics of facultatively symbiotic Frankia sp strains reflect host range and host plant biogeography. Genome research. 2007;17:7-15
24. Ngom M, Oshone R, Hurst SGt, Abebe-Akele F, Simpson S, Morris K. et al. Permanent Draft Genome Sequence for Frankia sp. Strain CeD, a Nitrogen-Fixing Actinobacterium Isolated from the Root Nodules of Casuarina equistifolia Grown in Senegal. Genome announcements. 2016:4
25. Oshone R, Ngom M, Abebe-Akele F, Simpson S, Morris K, Sy MO. et al. Permanent Draft Genome Sequence of Frankia sp. Strain Allo2, a Salt-Tolerant Nitrogen-Fixing Actinobacterium Isolated from the Root Nodules of Allocasuarina. Genome announcements. 2016:4
26. Hurst SGt, Oshone R, Ghodhbane-Gtari F, Morris K, Abebe-Akele F, Thomas WK. et al. Draft Genome Sequence of Frankia sp. Strain Thr, a Nitrogen-Fixing Actinobacterium Isolated from the Root Nodules of Casuarina cunninghamiana Grown in Egypt. Genome announcements. 2014/05/24 ed. 2014 p. e00493-14
27. Ghodhbane-Gtari F, Hurst SGt, Oshone R, Morris K, Abebe-Akele F, Thomas WK. et al. Draft Genome Sequence of Frankia sp. Strain BMG5.23, a Salt-Tolerant Nitrogen-Fixing Actinobacterium Isolated from the Root Nodules of Casuarina glauca Grown in Tunisia. Genome announcements. 2014;2:e00520-14
28. Mansour SR, Oshone R, Hurst SGt, Morris K, Thomas WK, Tisa LS. Draft Genome Sequence of Frankia sp. Strain CcI6, a Salt-Tolerant Nitrogen-Fixing Actinobacterium Isolated from the Root Nodule of Casuarina cunninghamiana. Genome announcements. 2014:2
29. D'Angelo T, Oshone R, Abebe-Akele F, Simpson S, Morris K, Thomas WK. et al. Permanent Draft Genome Sequence of Frankia sp. Strain BR, a Nitrogen-Fixing Actinobacterium Isolated from the Root Nodules of Casuarina equisetifolia. Genome announcements. 2016:4
30. Blin K, Medema MH, Kazempour D, Fischbach MA, Breitling R, Takano E. et al. antiSMASH 2.0-a versatile platform for genome mining of secondary metabolite producers. Nucleic Acids Res. 2013;41:W204-W12
31. Weber T, Blin K, Duddela S, Krug D, Kim HU, Bruccoleri R. et al. antiSMASH 3.0-a comprehensive resource for the genome mining of biosynthetic gene clusters. Nucleic Acids Res. 2015;43:W237-W43
Author contact
Corresponding authors: Louis S. Tisa Mailing address: Department of Molecular, Cellular, and Biomedical Sciences, University of New Hampshire, 46 College Rd., Durham, NH 03824-2617. Email: louis.tisaedu Telephone: 1-603-862-2442 Fax: 1-603-862-2621 and Hassen Gherbi Mailing address: LSTM-Laboratoire des Symbioses Tropicales & Méditerranéennes (IRD-CIRAD-INRA-UM-SupAgro), Campus International de Baillarguet, TA A-82/J. 34398 Montpellier Cedex 5 France. Email: hassen.gherbifr. Telephone: +33 (0)4 67 59 37 45. Fax: +33 (0)4 67 59 38 02

 Global reach, higher impact
Global reach, higher impact