ISSN: 1839-9940
J Genomics 2020; 8:43-48. doi:10.7150/jgen.39147 This volume Cite
Research Paper
A detailed characteristics of bias associated with long runs of homozygosity identification based on medium density SNP microarrays
1. University Centre of Veterinary Medicine, University of Agriculture in Kraków, Al. Mickiewicza 24/28, 30-059 Kraków, Poland.
2. National Research Institute of Animal Production, Department of Animal Molecular Biology, Krakowska 1, 32-083 Balice, Poland
3. College of Animal Science and Technology, Northwest A&F University, Yangling, Shaanxi 712100, China
Abstract
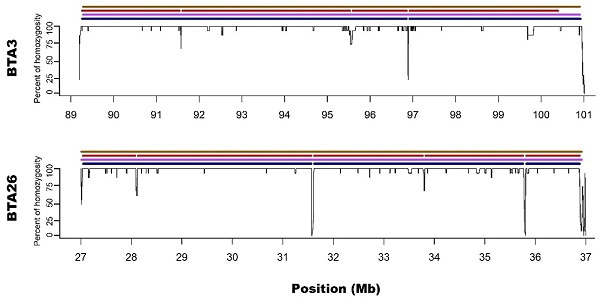
In the present study, runs of homozygosity (ROH) detected with the use of a standard bovine 54k single nucleotide polymorphism (SNP) genotyping assay and two different ROH detection approaches, based on 50 (M1) or 15 (M2) consecutive SNPs, were compared with results of whole genome sequencing. Both microarray-based methods accurately recognised medium-sized ROH, however, it was found that M2 method seemed to better than M1 identify short ROH, but highly overestimated their number, leading to numerous false positive calls. Moreover, long ROH identified with microarray data tended to break into shorter segments in sequencing data because of the presence of regions with high heterozygosity within the ROH sequences. This may indicate, that these long ROH are formed by closely positioned shorter homozygous segments that may be of older origin or may be created by two similar but not identical haplotypes, showing minor internal recombination signs. Such finding also suggests that at least some of the results of previous studies in regard to long ROH may be biased leading to inaccurate estimations of genomes autozygosity via ROH classification into length categories.
Keywords: runs of homozygosity, autozygosity, microarray, next generation sequencing

 Global reach, higher impact
Global reach, higher impact