ISSN: 1839-9940
J Genomics 2020; 8:84-88. doi:10.7150/jgen.51181 This volume Cite
Research Paper
Draft Genome Sequences for the Frankia sp. strains CgS1, CcI156 and CgMI4, Nitrogen-Fixing Bacteria Isolated from Casuarina sp. in Egypt
1. Faculty of Science, Suez Canal University, Ismailia, Egypt
2. University of New Hampshire, Durham, New Hampshire, USA
3. Present address: HM Clause, Davis, California, USA
Received 2020-7-27; Accepted 2020-9-5; Published 2020-9-23
Abstract
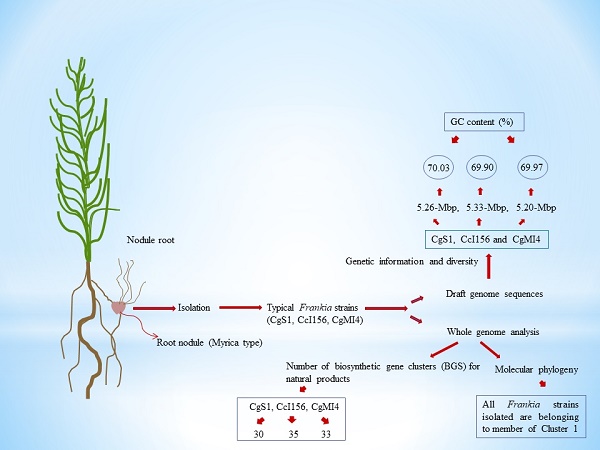
Frankia sp. strains CgS1, CcI156 and CgMI4 were isolated from Casuarina glauca and C. cunninghamiana nodules. Here, we report the 5.26-, 5.33- and 5.20-Mbp draft genome sequences of Frankia sp. strains CgS1, CcI156 and CgMI4, respectively. Analysis of the genome revealed the presence of high numbers of secondary metabolic biosynthetic gene clusters.
Keywords: actinobacteria, actinorhizal symbiosis, genomes, host microbe interactions, nitrogen fixation, natural products, salt tolerance
Introduction
Members of the genus Frankia are soil-dwelling Actinobacteria that form a symbiotic association with Angiosperms, representing eight different plant families [1-3]. This plant-microbe association, termed actinorhizal, aid these pioneer plants in colonizing harsh environments and disrupted lands [2, 4, 5]. Actinorhizal plants play a role economically in agroforestry, land reclamation, crop protection, and soil stabilization projects [4-7]. Based on molecular phylogeny, Frankia strains group into four major clusters that also follow plant host range specificity [8-11]. Cluster 1 consists of Frankia strains that associate with host plants in the Casuarinaceae, Betulaceae and Myricaceae families, while members of cluster 2 are infective on Rosaceae, Coriariaceae, Datiscaceae, and the genus Ceanothus (Rhamnaceae). Cluster 3 are the most promiscuous and are infective on Elaeagnaceae, Rhamnaceae, Myricaceae, Gynmnostoma, and occasionally the genus Alnus. Cluster 4 consists of “atypical” Frankia strains that are unable to re-infect actinorhizal host plants or form ineffective nonnitrogen-fixing root nodule structures.
Representative genomes from each cluster have been sequenced [12]. These Frankia genome databases has enabled the use of “omics” approaches [13-15] and allowed speciation of the genus [16].
Cluster 1 is further divided into subclades with subclade Ic containing strains limited to Casuarina and Allocasuarina and Myricaceae host plants. Actinorhizal plants belonging to Casuarinaceae family such as Casuarina glauca and C. equisetifolia will grow under saline environments [4, 5, 13]. These fast-growing trees originated from Australia and Pacific islands, and are widely used in agroforestry systems for several purposes [17]. Large-scale planting of casuarinas has proven to have a strong impact especially in China, Senegal, Egypt and Tunisia [18]. In arid and semi-arid areas, salinization of soils and groundwater is a serious problem causing a drastic reduction in agricultural production ([19, 20]; http://www.fao.org/nr/land/degradation/en/). Over 800 million hectares of land throughout the world are salt-affected [20]. An effective method for the reclamation of salt-affected soils involves initiating plant succession using fast growing, nitrogen fixing actinorhizal trees such as the Casuarina. The salt tolerance of Casuarina is enhanced by the nitrogen-fixing symbiosis that they form with Frankia.[7]. A broad spectrum “omics” approach on Frankia strains from Casuarina plants has allowed us to define several genes involved salt tolerance [21] including one gene that was cloned into a salt-sensitive strain [22]. The expression of the cloned genes provided increased tolerance to salt in the receipt strain. Here we have isolated three Frankia strains from Casuarina species that were found under salt stress conditions. These strains were sequenced to provide more information about salt tolerance genes and a greater understanding the metabolic potential of these actinobacteria.
Isolation of Frankia strains
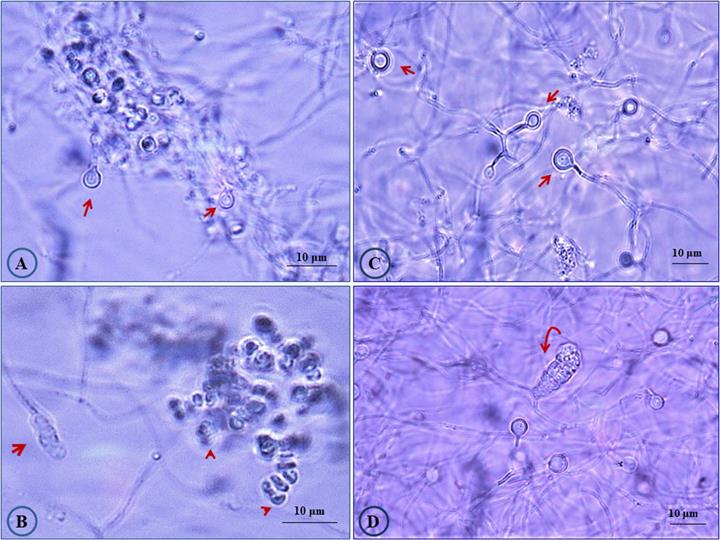
Three different Casuarina plantation sites in Egypt were chosen for sampling and root nodules were collected and transferred to the laboratory in an icebox. From site one, the nodule sample was collected from a large Casuarina glauca. tree growing in sandy loamy soil around a Mango Garden, as “green-barrier fence, in Abuo-Shata, Ismailia, Egypt. For the second site, nodules were collected from Casuarina cunninghamiana tree growing inside a nearby Mango garden in Abuo-Shata, Ismailia, Egypt (Latitude and longitude coordinates: 30°35′47.37″N, 32°16′17.25″E). The third nodule sample was collected from C. glauca growing in sandy soil along Ismailia-Cairo highway (30.0444° N, 31.2357° E). After the nodules were cleaned with tap water and surface sterilized, Frankia strains were isolated as previously described [23]. Frankia isolates obtained from site 1, 2 and 3 were designated CgIM4, CcI156 and CgIS1, respectively, and propagated in B medium [23]. The Frankia isolates displayed the typical morphological traits of the genus Frankia (Fig. 1). These developmentally complex bacteria form three cell types: vegetative hyphae, spores located in sporangia and the unique lipid-enveloped cellular structures, termed vesicles [1, 3]. Vesicles act as specialized structures for the nitrogen fixation process [1, 3].
Photomicrograph showing morphological features of Frankia sp. strains CgIM4 and CcI156. Cultures were grown in liquid B medium [23] for 45 days. (A) Frankia sp. strain CgIM4 showing typical active vesicles with double wall (arrow); (B) sporulation of Frankia sp. strain CgIM4 forming typical multilocular sporangium (arrow) and the release of mature spores (head arrows); (C) healthy vesicles (arrows) for Frankia sp. strain CcI156; (D) Frankia sp. strain CcI156 intercalary multilocular sporangium with acropetal spore maturation (arrow).
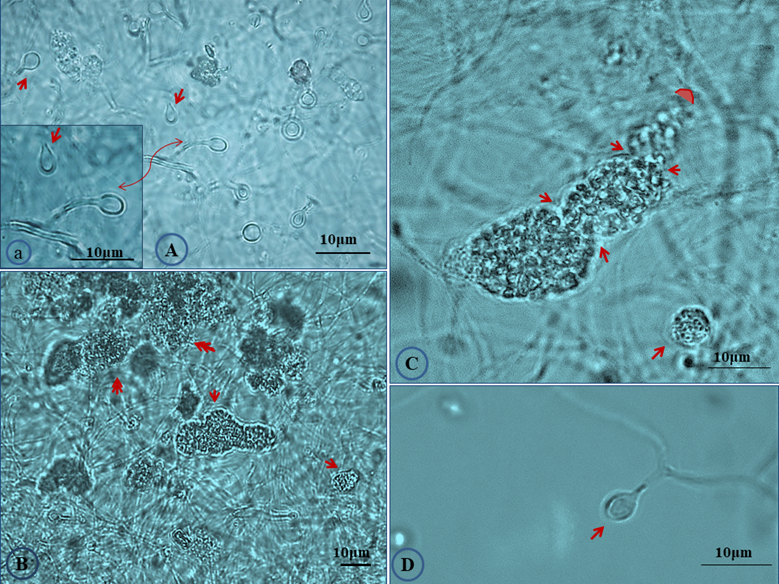
The salt tolerance levels of Frankia sp. strains CcI156 and CgMI4 were determined by measuring the total cellular protein content after growth under salt stress, as described previously [21]. Frankia sp. strain CcI6, a salt-tolerant strain, and Frankia casuarinae strain CcI3, a salt-sensitive strain, were included as controls (Fig. S1). Frankia sp. strains CcI156 and CgMI4 have MIC values of 550 and 700 mM NaCl, respectively. Frankia sp. strain CcI6 and F. casuarinae strain CcI3 have MIC values of 1,000 and 475 mM NaCl, respectively, similar to previous results [21]. The MIC value for Frankia sp. strains CcI156 was similar to the values obtained for other Frankia lineage Ic strain (Frankia sp. strains BR, BMG5.23, CeD and Thr), while Frankia sp. strain CgMI4 was greater than those strains but not to the level of Frankia sp. strains CcI6 and Allo2 [21]. Frankia sp. strains CgMI4 and CcI156 exhibited several morphological changes when grown under salt-stress conditions (Fig. 2; Fig S2). Both sporangium and vesicle development were altered.
Effect of salt stress on the morphological characters of Frankia sp. strain CgIM4. Cultures were grown in B medium [23] under 300 mM NaCl stress for 45 days. (A) Vesicles showed slight deformation in which the elliptical shape with thinner wall was observed (arrow) compared to thick circular shape in normal condition; (a) magnified section of panel A showing deformed vesicles; (B) abundance of mature sporangia (arrows) were detected at a high frequency; (C) twisted mature sporangia (arrows) with constriction which showed mature spores filling the whole sporangium, note the absence of acropetal spore maturation as under non-stressed conditions (Fig. 1); (D) deformed vesicle.
Sequencing of Frankia strains CgS1, CcI156 and CgMI4
Sequencing of the draft genomes of Frankia sp. strains CgS1, CcI156 and CgMI4 was performed at the Hubbard Center for Genome Studies (University of New Hampshire, Durham, NH) using Illumina technology techniques [24]. For each genome, a standard Illumina shotgun library was constructed and sequenced using the Illumina HiSeq2500 platform, using a pair-end library with an average size of 600 bp obtaining 5,345,232, 11,011,806 and 11,746,204 reads of 250 bp in length for strains CgS1, CcI156 and CgMI4, respectively (Table 1). The Illumina sequence data were trimmed by Trimmonatic version 0.32 [25], assembled using Spades version 3.5 [25] and ALLPaths-LG version r52488 [26]. The metrics for final draft assemblies for Frankia sp. strains CgS1, CcI156 and CgMI4 are found in Table 1. The two 135 and 145 contigs ranged from 5.2 to 5.3 Mbp in size with 70% G + C content.
Genome features of Frankia sp. strain CgIS1, CcI156 and CgMI4 isolated from Casuarina root nodules.
| Strain | CgIS1 | CcI156 | CgMI4 |
|---|---|---|---|
| Reads | 5,345,232 | 11,011,806 | 11,746,204 |
| N50 (kb) | 99.0 | 80.6 | 90.5 |
| Contigs | 135 | 145 | 135 |
| Genome size | 5,257,145 | 5,330,592 | 5,199,090 |
| Coverage | 157.9X | 427.0X | 328.1X |
| G + C content (%) | 70.03 | 69.90 | 69.97 |
| CDS (coding) | 4,224 | 4,304 | 4,165 |
| rRNA | 2 | 2 | 0 |
| tRNA | 45 | 45 | 46 |
| GeneBank Accession number | MBLP00000000 | MOMD00000000 | MAQZ00000000 |
| BioProject | PRJNA299691 | PRJNA299689 | PRJNA286879 |
The assembled genomes were annotated via the NCBI Prokaryotic Genome Annotation Pipeline (PGAP) [27]. The annotation features are also given in Table 1. The genome features of three Frankia sp. strains are similar to other cluster 1c genomes (Table 1) including F. casuarinae strain CcI3T [28]. Analysis of the three genomes for the number of genes associated with the Clusters of Orthologous Groups (COG) functional categories showed that the pattern of distribution was similar to F. casuarinae strain CcI3T (Table S1). Phylogenetic analysis of the 16S rDNA shows that Frankia sp. strains CgS1, CcI156 and CgMI4 group with the cluster 1c strains (Figure S3) and further confirmed by dendrogram of the entire genomes (Figure S4). A whole genome-based taxonomic analysis via the Type (Strain) Genome Server (TYGS) platform [29] (https://tygs.dsmz.de) including digital DNA:DNA hybridization (dDDH) values [30] was performed to identify these strain compared to other Frankia genomes from Cluster 1c (Table S2). The type-based species clustering using a 70% dDDH radius around each of the type strains was used as previously [31], while subspecies clustering was done using a 79% dDDH threshold as previously introduced [32]. These data indicate that Frankia sp. strains CgS1, CcI156 and CgMI4 are members of Frankia casuarinae species. Average nucleotide identity (ANI) analysis of these genomes (Table S3) confirmed that idea with an ANI threshold range of 95% for species delineation [33].
Bioinformatic analysis of this genomes by the use of the AntiSMASH program [34] revealed the presence of high numbers of secondary metabolic biosynthetic gene clusters (Table 2), which is consistent with previous results with other Frankia lineage 3 genomes including cluster 3 [12, 35]. Many of these potential natural products should be involved in the plant-microbe interactions and aiding in their plant-growth-promoting activities. Analysis of the Frankia sp. strain CgS1, CcI156 and CgMI4 genomes revealed similar salt-tolerance genes as previously identified including several hypothetical proteins responsive to salt stress detected by genomic, transcriptomic, and proteomic approaches [21].
Biosynthetic gene clusters (BGS) for natural products found in the select Frankia genomes
| Strain | CgS1 | CcI156 | CgMI4 | CcI6 | CcI3 |
|---|---|---|---|---|---|
| Total BGC Clusters | 30 | 35 | 33 | 33 | 29 |
| NRPS1 | 6 | 9 | 7 | 8 | 3 |
| PKS2 | 5 | 8 | 8 | 8 | 5 |
| Terpene | 4 | 4 | 4 | 4 | 4 |
| Siderophore | 1 | 1 | 1 | 1 | 1 |
| Bacteriocin Lassopeptide | 2 | 2 | 2 | 3 | 3 |
| Lantipeptide | 5 | 5 | 6 | 5 | 6 |
| Reference | This study | This study | This study | [36] | [37] |
Biosynthettic gene clusters were identified by the use of the AntiSMASH software [34]. 1 NRPS: Nonribosomal peptide synthase. 2 PKS: polyketide synthase including Type I, II, III, Trans-AT and other types
In summary, the Frankia sp. strain CgS1, CcI156 and CgMI4 genome has revealed an interesting metabolic potential including secondary metabolites pathways and natural product profiles and serves as more representatives of Frankia cluster 1c.
Nucleotide sequence accession numbers
These whole-genome shotgun sequences have been deposited at DDBJ/EMBL/GenBank under the accession numbers: MBLP00000000, MOMD00000000 and MAQZ00000000 for strains CgS1, CcI156 and CgMI4, respectively. The versions described in this paper are the first versions, MBLP 01000000, MOMD01000000, and MAQZ01000000 for strains CgS1, CcI156 and CgMI4, respectively. Both the assembly and raw reads are available at DDBJ/ENA/GenBank under BioProject number PRJNA299691, PRJNA299689, and PRJNA286879 for strains CgS1, CcI156 and CgMI4, respectively.
Supplementary Material
Supplementary figures and tables.
Acknowledgements
Partial funding was provided by the New Hampshire Agricultural Experiment Station. This is Scientific Contribution Number 2863. This work was also supported by the USDA National Institute of Food and Agriculture Hatch 022821 (LST), the Plant Health and Production and Plant Products Program grant number 2015-67014-22849/project accession no. 1005242 from the USDA National Institute of Food and Agriculture (LST), the College of Life Science and Agriculture at the University of New Hampshire-Durham, and Technology Development Fund (STDF), grant No. 900, as International Cooperation Project (SM and LST). Sequencing was performed on an Illumina HiSeq2500 purchased with an NSF MRI Grant: DBI-1229361 to WK Thomas.
Competing Interests
The authors have declared that no competing interest exists.
References
1. Normand P, Benson DR, Berry AM, Tisa LS. Family Frankiaceae. In: Rosenberg E, DeLong EF, Lory S, Stackebrandt E, Thompson F, editors. The Prokaryote - Actinobacteria Berlin Heidelberg: Springer-Verlag. 2014 p. 339-56
2. Benson DR, Dawson JO. Recent advances in the biogeography and genecology of symbiotic Frankia and its host plants. Physiol Plantarum. 2007;130:318-30
3. Benson DR, Silvester WB. Biology of Frankia Strains, Actinomycete Symbionts of Actinorhizal Plants. Microbiol Rev. 1993;57:293-319
4. Diagne N, Arumugam K, Ngom M, Nambiar-Veetil M, Franche C, Narayanan KK. et al. Use of Frankia and Actinorhizal Plants for Degraded Lands Reclamation. Biomed Res Int. 2013: e948258.
5. Ngom M, Oshone R, Diagne N, Cissoko M, Svistoonoff S, Tisa LS. et al. Tolerance to environmental stress by the nitrogen-fixing actinobacterium Frankia and its role in actinorhizal plants adaptation. Symbiosis. 2016;70:17-29
6. Diagne N, Ngom M, Djighaly PI, Ngom D, Ndour B, Cissokho M. et al. Remediation of heavy-metal-contaminated soils and enhancement of their fertility with actinorhizal plants. In: Sherameti I, Varma A, editors. Heavy Metal Contamination of Soils, Soil Biology. Switzerland: Springer International Publishing. 2015 p. 355-66
7. Ngom M, Gray K, Diagne N, Oshone R, Fardoux J, Gherbi H. et al. Symbiotic Performance of Diverse Frankia Strains on Salt-Stressed Casuarina glauca and Casuarina equisetifolia Plants. Frontiers in Plant Science. 2016 p. 1331
8. Normand P, Orso S, Cournoyer B, Jeannin P, Chapelon C, Dawson J. et al. Molecular phylogeny of the genus Frankia and related genera and emendation of the family Frankiaceae. Int J Syst Bacteriol. 1996;46:1-9
9. Gtari M, Tisa LS, Normand P. Diversity of Frankia Strains, Actinobacterial Symbionts of Actinorhizal Plants. In: Aroca R, editor. Symbiotic Endophytes. 2013 p. 123-48
10. Nouioui I, Ghodhbane-Gtari F, Beauchemin NJ, Tisa LS, Gtari M. Phylogeny of members of the Frankia genus based on gyrB, nifH and glnII sequences. Anton Leeuw Int J G. 2011;100:579-87
11. Ghodhbane-Gtari F, Nouioui I, Chair M, Boudabous A, Gtari M. 16S-23S rRNA Intergenic Spacer Region Variability in the Genus Frankia. Microb Ecol. 2010;60:487-95
12. Tisa LS, Oshone R, Sarkar I, Ktari A, Sen A, Gtari M. Genomic approaches toward understanding the actinorhizal symbiosis: an update on the status of the Frankia genomes. Symbiosis. 2016;70:5-16
13. Diagne N, Djighaly PI, Ngom M, Pesce C, Champion A, Svistoonoff S, et al. Advances in Frankia genome studies and molecular aspects of tolerance to environmental stresses.. In: Salwan R, Sharma V, Yang W, Khalil L, Al-Ani T, editors. Molecular Aspects of Plant Beneficial Microbes in Agriculture. Cambridge, MA, USA: Elsevier; 2020. p. 381-90
14. Nouioui I, Cortes-Albayay C, Carro L, Castro JF, Gtari M, Ghodhbane-Gtari F. et al. Genomic Insights Into Plant-Growth-Promoting Potentialities of the Genus Frankia. Front Microbiol. 2019;10:1457
15. Gherbi H, Hocher V, Ngom M, Diagne N, Fournier J, Carre-Mlouka A. et al. Molecular methods for research on actinorhiza. In: Reinhardt D, Sharna AK, editors. Molecular Aspects of Plant Beneficial Microbes in Agriculture. Cambridge, MA, USA Elsevier. 2019 p. 381-90
16. Gtari M, Nouioui I, Sarkar I, Ghodhbane-Gtari F, Tisa LS, Sen A. et al. An update on the taxonomy of the genus Frankia Brunchorst, 1886, 174(AL). Antonie Van Leeuwenhoek International Journal of General and Molecular Microbiology. 2019;112:5-21
17. Dommergues YR. Contribution of actinorhizal plants to tropical soil productivity and rehabilitation. Soil Biol Biochem. 1997;29:931-41
18. Gtari M, Dawson JO. An overview of actinorhizal plants in Africa. Functional Plant Biology. 2011;38:653-61
19. Munns R. Genes and salt tolerance: bringing them together. New Phytologist. 2005;167:645-63
20. Rengasamy P. World salinization with emphasis on Australia. J Exp Bot. 2006;57:1017-23
21. Oshone R, Ngom M, Chu F, Mansour S, Sy MO, Champion A. et al. Genomic, transcriptomic, and proteomic approaches towards understanding the molecular mechanisms of salt tolerance in Frankia strains isolated from Casuarina trees. BMC Genomics. 2017;18:633
22. Pesce C, Oshone R, Hurst SGt, Kleiner VA, Tisa LS. Stable Transformation of the Actinobacteria Frankia spp. Appl Environ Microbiol. 2019 85
23. Mansour SR, Dewedar A, Torrey JG. Isolation, Culture, and Behavior of Frankia Strain HFPCgI4 from Root Nodules of Casuarina glauca. Bot Gaz. 1990;151:490-6
24. Bennett S. Solexa Ltd. Pharmacogenomics. 2004;5:433-8
25. Bolger AM, Lohse M, Usadel B. Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics. 2014;30:2114-20
26. Gnerre S, MacCallum I, Przybylski D, Ribeiro FJ, Burton JN, Walker BJ. et al. High-quality draft assemblies of mammalian genomes from massively parallel sequence data. P Natl Acad Sci USA. 2011;108:1513-8
27. Tatusova T, DiCuccio M, Badretdin A, Chetvernin V, Nawrocki EP, Zaslavsky L. et al. NCBI prokaryotic genome annotation pipeline. Nucleic Acids Res. 2016;44:6614-24
28. Nouioui I, Ghodhbane-Gtari F, Montero-Calasanz MD, Goker M, Meier-Kolthoff JP, Schumann P. et al. Proposal of a type strain for Frankia alni (Woronin 1866) Von Tubeuf 1895, emended description of Frankia alni, and recognition of Frankia casuarinae sp. nov. and Frankia elaeagni sp. nov. Int J Syst Evol Microbiol. 2016;66:5201-10
29. Meier-Kolthoff JP, Goker M. TYGS is an automated high-throughput platform for state-of-the-art genome-based taxonomy. Nature communications. 2019;10:2182
30. Meier-Kolthoff JP, Auch AF, Klenk HP, Goker M. Genome sequence-based species delimitation with confidence intervals and improved distance functions. Bmc Bioinformatics. 2013;14:Artn 60
31. Liu Y, Lai Q, Goker M, Meier-Kolthoff JP, Wang M, Sun Y. et al. Genomic insights into the taxonomic status of the Bacillus cereus group. Sci Rep. 2015;5:14082
32. Meier-Kolthoff JP, Hahnke RL, Petersen J, Scheuner C, Michael V, Fiebig A. et al. Complete genome sequence of DSM 30083(T), the type strain (U5/41(T)) of Escherichia coli, and a proposal for delineating subspecies in microbial taxonomy. Stand Genomic Sci. 2014 9
33. Goris J, Konstantinidis KT, Klappenbach JA, Coenye T, Vandamme P, Tiedje JM. DNA-DNA hybridization values and their relationship to whole-genome sequence similarities. Int J Syst Evol Micr. 2007;57:81-91
34. Blin K, Shaw S, Steinke K, Villebro R, Ziemert N, Lee SY. et al. antiSMASH 5.0: updates to the secondary metabolite genome mining pipeline. Nucleic Acids Res. 2019;47:W81-W7
35. Udwary DW, Gontang EA, Jones AC, Jones CS, Schultz AW, Winter JM. et al. Significant Natural Product Biosynthetic Potential of Actinorhizal Symbionts of the Genus Frankia, as Revealed by Comparative Genomic and Proteomic Analyses. Appl Environ Microb. 2011;77:3617-25
36. Mansour SR, Oshone R, Hurst SGt, Morris K, Thomas WK, Tisa LS. Draft Genome Sequence of Frankia sp. Strain CcI6, a Salt-Tolerant Nitrogen-Fixing Actinobacterium Isolated from the Root Nodule of Casuarina cunninghamiana. Genome announcements. 2014;2:e01205-13
37. Normand P, Lapierre P, Tisa LS, Gogarten JP, Alloisio N, Bagnarol E. et al. Genome characteristics of facultatively symbiotic Frankia sp strains reflect host range and host plant biogeography. Genome research. 2007;17:7-15
Author contact
Corresponding authors: Louis S. Tisa. Mailing address: Department of Molecular, Cellular, and Biomedical Sciences, University of New Hampshire, 46 College Rd., Durham, NH 03824-2617. Email: louis.tisaedu; Telephone: 1-603-862-2442; Fax: 1-603-862-2621. Samira Mansour. Mailing address: Faculty of Science, Suez Canal University, Ismailia, Egypt. Email: samirarmansourcom

 Global reach, higher impact
Global reach, higher impact