ISSN: 1839-9940
J Genomics 2015; 3:40-50. doi:10.7150/jgen.11015 This volume Cite
Review
The Multifunctions of WD40 Proteins in Genome Integrity and Cell Cycle Progression
1. Department of Biochemistry and Molecular Genetics, University of Colorado School of Medicine, Aurora, CO, 80045, USA
2. Orthopedics Department, Changhai Hospital Affiliated to Second Military Medical University, Shanghai, 200433, China
Published 2015-2-5
Abstract
Eukaryotic genome encodes numerous WD40 repeat proteins, which generally function as platforms of protein-protein interactions and are involved in numerous biological process, such as signal transduction, gene transcriptional regulation, protein modifications, cytoskeleton assembly, vesicular trafficking, DNA damage and repair, cell death and cell cycle progression. Among these diverse functions, genome integrity maintenance and cell cycle progression are extremely important as deregulation of them is clinically linked to uncontrolled proliferative diseases such as cancer. Thus, we mainly summarize and discuss the recent understanding of WD40 proteins and their molecular mechanisms linked to genome stability and cell cycle progression in this review, thereby demonstrating their pervasiveness and importance in cellular networks.
Keywords: WD40 protein, DNA damage, cell death, cell cycle
1. Introduction: Overviews of WD40 proteins and their functions
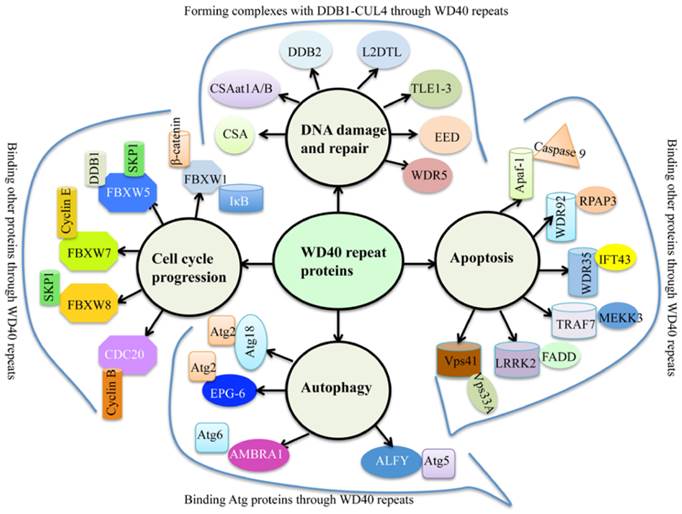
WD40 repeats proteins are abundant in eukaryotes, but rarely present in prokaryotes [1]. Human genome encodes 349 predicted WD40 repeat-containing proteins [2], and plant genomes harbor more than 200 putative WD40 proteins [1, 3, 4]. Generally, WD40 proteins are characterized by the presence of repeating units with 44-60 variable residues that ended with tryptophan (W) and aspartate (D) dipeptides [5]. The basic function of WD40 proteins is to serve as rigid platforms for protein-protein and protein-DNA interactions [6]. During the past decades, multiple WD40 protein complexes have been identified, leading to a better understanding of WD40 domain functions involved in many biological processes such as signal transduction, gene expression regulation, protein ubiquitination and histone methylation, genome stability and cell cycle control [1, 6]. Interestingly, a variety of proteins contain conserved “WDxR” motif(s) in their WD40 repeat regions, such as CSA (Cockayne syndrome A) [7], CSAat1A/B (CSA-like proteins in Arabidopsis 1A and 1B) [8], DDB2 (Damaged DNA binding protein 2) [9], L2DTL (Drosophila lethal(2) denticleless protein) [10], TLE1-3 (Transducin-like enhancer of split 1 to 3) [11], EED (Embryonic ectoderm development protein) [11], and WDR5 (WD repeat-containing protein 5) [11] (Figure 1). These proteins commonly form complexes with DDB1 (damaged DNA binding protein 1) through the “WDxR” motif in response to DNA damage and repair (Figure 1). In addition, numerous WD40 proteins also have been identified to interact with other proteins in response to apoptosis, autophagy and cell cycle progression (Figure 1). These WD40 proteins mainly include Apaf-1 (Activation of apoptotic protease activating factor 1) [12], WDR35 [13], WDR92 [14], TRAF7 (Tumor necrosis factor receptor-associated factor) [15], LRRK2 (Leucine-rich repeat kinase 2) [16], Vps41 (Vacuolar protein sorting 41) [17], Atg18 (Autophagy-related protein 18) [18], EPG-6 (Ectopic PGL granules 6) [19], AMBRA1 (Autophagy/beclin-1 regulator 1) [20], ALFY (autophagy‐linked FYVE protein) [21], FBXW1 (F-box/WD repeat-containing protein 1) [22], FBXW5 [23], FBXW7 [24], FBXW8 [25], and CDC20 (Cell division cycle protein 20) [26].
The involvement of WD40 proteins in genome stability and cell cycle progression. Multiple WD40 proteins interact with DDB1, Atg proteins, Cyclin proteins and other proteins through their WD40 repeats in response to DNA damage and repair, autophagy, cell cycle progression and/or apoptosis. CSA, Cockayne syndrome A; CSAat1A/B, CSA-like proteins in Arabidopsis 1A and 1B; DDB2, Damaged DNA binding protein 2; L2DTL, Drosophila lethal(2) denticleless protein; TLE1-3, Transducin-like enhancer of split 1 to 3; EED, Embryonic ectoderm development protein; WDR5, WD40 repeat-containing protein 5; Apaf-1, Activation of apoptotic protease activating factor 1; WDR35/92, WD40 repeat-containing protein 35/92; TRAF7, Tumor necrosis factor receptor-associated factor; LRRK2, Leucine-rich repeat kinase 2; Vps41, Vacuolar protein sorting 41; Atg18, Autophagy-related protein 18; EPG-6, Ectopic PGL granules 6; AMBRA1, autophagy/beclin-1 regulator 1; ALFY, autophagy‐linked FYVE protein; FBXW1/5/7/8, F-box/WD repeat-containing protein 1, 5, 7 and 8; CDC20,Cell division cycle protein 20.
Some of these WD40 proteins involved in genome stability and cell cycle progression have been determined their structures (Table 1). Generally, the WD40 domain exhibits a β-propeller architecture, which typically contains seven blades and each of them is composed of four anti-parallel β-sheets [1]. WD40 domain recognizes and binds to other proteins mainly through the smaller top surface, but also through the bottom surface and sides in some cases [1]. Clinically, inherited or acquired defects of WD40 proteins always lead to severe health problems and are genetically associated with a variety of diseases such as Cockayne syndrome [27], multiple cancer types [28], Lissencephaly [29] and Parkinson's disease [30].
The bind partners, binding domain/motifs, biological functions and available structures of WD40 proteins
| WD40 protein | Binding partner | Binding domain and consensus motif | Biological functions | Available structure | References |
|---|---|---|---|---|---|
| CSA | DDB1, RNA polymerase II TFIIH | WD40 domain, VKLWDVRR | TCR DNA repair | Yes | [47] |
| CSAat1A/1B | DDB1A | WD40 domain, DGAIRFWDIR | TCR DNA repair | No | [8] |
| DDB2 | DDB1, E2F1 | WD40 domain, VKIWDLR | GGR DNA repair | Yes | [47] |
| WDR5 | DDB1 Histone H3 | WD40 domain, R-X-Kdimethylation | Chromatin regulator | Yes | [11] |
| L2DTL (CDT2) | DDB1, CDT1 | WD40 domain, IKVWDLRK | DNA dmage, cell cycle control | Yes | [10] |
| EED | DDB1 | WD40 domain, LRLWNIQTD | Chromatin regulator | Yes | [121] |
| WDR18 | TopBP1 | WD40 domain, Unknown motif | DNA damage checkpoint | No | [56] |
| TLE1-3 | DDB1 | WD40 domain, VRSWDLREGR | Transcription regulator | Yes | [122] |
| Apaf-1 | Caspase 9 | WD40 domain Unknown motif | Apoptosis | Yes | [123] |
| WDR92 | RPAP3 | WD40 domain, Unknown motif | Apoptosis | Yes | [1] |
| WDR35 | IFT43, BCL6 | WD40 domain, Unknown motif | Apoptosis | Predicted structure | [124] |
| TRAF7 | MEKK3 | WD40 domain, Unknown motif | Apoptosis | No | [15] |
| Vps41 | VPS33A, VPS16, 18 | WD40 domain, Unknown motif | Apoptosis | No | [17] |
| VprBP | DDB1 | WD40 domain, VLWDVRSAQ | Apoptosis, cell cycle control | Yes | [125] |
| LRRK2 | FADD | WD40 domain, Unknown motif | Apoptosis | Yes | [76] |
| RACK1 | p63 | WD40 domain, Unknown motif | Apoptosis | Yes | [78] |
| Atg18 | Atg2 | WD40 domain IYQFRRGTYA | Autophagy | Yes | [126] |
| EPG-6 | Atg2 | WD40 domain Unknown motif | Autophagy | No | [19] |
| AMBRA1 | Atg6, PIK3C3 | WD40 domain Unknown motif | Autophagy | No | [20] |
| ALFY | Atg5, GABARAP | WD40 domain DGFIFV | Autophagy | Yes | [21] |
| LYST | LAST1, LAST2 | Unkown domain and motif | Autophagy | No | [101] |
| LRBA | WDR23 | Unkown domain and motif | Autophagy | No | [102] |
| WDR26 | DDB1 | Unkown domain and motif | Cell death | No | [103] |
| MDV1 | Dnm1 | WD40 domain, Unknown motif | Cell death | No | [104] |
| FBXW1 | β-catenin, IκB | WD40 domain, D-pS-G-Φ-X-pS | Cell cycle control | Yes | [127] |
| FBXW5 | SKP1, DDB1 | WD40 domain, IWDRHY | DNA damage and cell cycle control | No | [128] |
| FBXW7 | Cyclin E, Sic1p, Gcn4p | WD40 domain, L-I/L/P-pT-P | Cell cycle control | Yes | [129] |
| FBXW8 | SKP1 | WD40 domain, Unokwn motif | Cell proliferation | No | [25] |
| CDC20 | Cyclin B | WD40 domain, Unknown motif | Cell cycle control | Yes | [112] |
2. WD40 proteins, DNA damage and repair
The genotoxic chemicals, ionizing radiation (IR) and ultraviolet (UV) radiation can cause DNA damage [31, 32]. Organisms have evolved complex and highly conserved systems to rapidly and efficiently repair damaged DNA. Commonly, DNA double-strand breaks (DSBs) are repaired through three distinct pathways, including non-homologous end-joining (NHEJ), alternative NHEJ and homologous recombination (HR) [31, 33]. UV-induced DNA lesions and other DNA bulks are removed from duplex DNA by nucleotide excision repair (NER) [31, 34]. Small and non-helix-distorting base lesions that affect only one DNA strand are repaired by base-excision repair (BER) [31]. Erroneous insertion, deletion and misincorporation of DNA bases are recognized and repaired by mismatch repair (MMR) [31]. DNA crosslinks, including intrastrand and interstrand crosslink, are repaired by Fanconi anemia (FA) pathway [31].
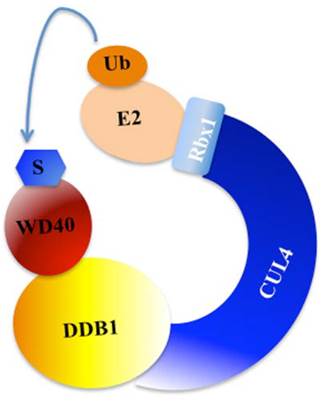
Post-translational modifications, such as phosphorylation, ubiquitination and sumoylation, play crucial roles in coordinating cellular response to DNA damage [31]. Previous studies have indicated that ubiquitination regulates and coordinates pathways involved in DNA damage recognition, signaling and repair [31, 35]. The eukaryotic ubiquitin-proteasome system (UPS) involves three enzymes: the E1 Ub-activating enzyme, E2 Ub-conjugating enzyme, and E3 Ub-protein ligase [36]. In mammals, cullin family proteins associate with RING proteins, adaptor proteins and substrate recognition receptors, forming the largest family of E3 ubiquitin ligases, namely, Cullin-RING ubiquitin ligases (CRLs) [37, 38]. Among these CRLs, CRL4 has been widely reported to function in the regulation of DNA damage and repair processes [37, 38]. CRL4 E3 ligase is consist of cullin 4A or 4B (CUL4A or CUL4B), RING-box protein 1 (RBX1, also known as ROC1), adaptor protein DDB1, and substrate recognition receptor DCAFs (DDB1 and CUL4-associated factors, also referred to as WD40 proteins) [37, 38]. Interestingly, the specificity of CRL4 E3 ligases is conferred by these WD40-containing substrate receptors [37, 38]. A variety of WD40 proteins, such as CSA, CSAat1A/1B, DDB2, WDR5, TLE1-3, L2DTL and EED, have been identified to interact with CUL4A/B-DDB1 complex in response to DNA damage (Figure 2) [7, 39].
A model for the assembly of the CUL4-DDB1-WD40 ubiquitin ligase complex. CUL4 serves as a rigid scaffold, and directly contacts with DDB1 with its N-terminal domain. CUL4 can also recruit Rbx1 and the E2 ubiquitin-conjugating enzyme via its C-terminal domain. The DDB1 further binds a variety of WD40 proteins to form E3 ligases, recognizing different substrates in response to different cellular processes [9]. S, substrate. Ub, ubiquitination.
Cockayne syndrome (CS) is a genetic recessive disorder characterized by growth retardation, impairment of nervous system development, hypersensitivity to sunlight and premature aging [40, 41]. CS can be resulted from mutations in either CSA (also known as ERCC8) or CSB gene (also known as ERCC6) [42]. Previous studies have demonstrated that CSA and CSB act in a transcription-coupled repair (TCR) pathway [43]. CSA is a subunit of CRL4 E3 ligase complex and CSB is a substrate of this E3 ligase [43]. After DNA damage caused by UV irradiation, CSB is ubiquitinated by the CRL4CSA E3 ligase and degraded by the proteasome at a late stage of the repair process, and that CSB degradation affects the recovery of RNA synthesis after TCR [43]. Interestingly, CSA contains two WDxR motifs in WD40 repeats and the second WDxR motif plays critical role for its interaction with DDB1 [7]. Further studies have revealed that CSAR217A mutation abolishes its binding to DDB1 in vitro, but not CSAR218A mutation [7]. Additionally, CSAA205P mutation, which is also located in the second WDxR motif, is identified to cause CS and block DDB1 binding [7]. In Arabidopsis, two CSA-like proteins, namely, CSAat1A and CSAat1B can form an E3 ligase with CUL4 and DDB1A to regulate substrates in response to DNA damage through TCR [8]. Both CSAat1A and 1B contain a WDxR motif in the WD40 domain, and single mutations at positions Asp-212, Trp-218, Asp-219, or Arg-221 in CSAat1A and 1B reduce their ability to interact with DDB1A [8].
The CUL4-DDB1 complex specifically binds to DDB2 to form the UV-damaged DNA-binding protein complex (the UV-DDB complex) [44], which further recognizes UV-induced DNA damage and recruits proteins of the NER pathway to initiate DNA repair [44, 45]. The UV-DDB complex preferentially binds to cyclobutane pyrimidine dimers (CPD), 6-4 photoproducts (6-4 PP), apurinic sites and short mismatches [46]. Biochemically, the CRL4DDB2 E3 ligase serve functions in global genome repair (GGR) after UV irradiation [47]. Similarly, DDB2 also contains a WDxR motif in the WD40 domain, which is critically important for DDB1-DDB2 interaction [45]. The R273H mutation within the WDxR motif is deficient for DDB1 binding and is causally associated with human xeroderma pigmentosum (XP-E) [45]. In human cells, the CRL4DDB2 E3 ligase can ubiquitinate histone H3, H4, and XP-C (xeroderma pigmentosum group C) protein to regulate their activities [48].
The immunoprecipitation and mass spectrometry assays have identified a variety of other WD40 proteins associated with CUL4-DDB1 complex, such as WDR5, TLE1-3 (TLEs), L2DTL and EED [49]. WDR5 is a core subunit of the human MLL (mixed-lineage leukaemia) and SET1 (hCOMPASS) histone H3 Lys4 (H3K4) methyltransferase complex [50]. TLEs are transcriptional corepressors that bind to a number of transcription factors, thereby inhibiting NF-kappa-B-regulated gene expression, and inhibiting transcriptional activation mediated by FOXA2 (forkhead box protein A2) [51], by CTNNB1 (catenin beta-1) and TCF (transcription factor) family members in Wnt signaling [52, 53]. The crystal structures of WD40-repeat domains in WDR5 and TLE1 indicate that the Asp and Arg/Lys in the WDxR/K motif are exposed on the bottom surface of the barrel-shaped WD40 propeller folds, and are thus available for interaction with CUL4-DDB1 [49]. L2DTL also requires the WD40-repeat domain for DDB1 binding and it is responsible for CDT1 proteolysis in response to DNA damage [49]. The CRL4L2DTL E3 ligase can promote the ubiquitination of proteins in S phase and after DNA damage [49]. Numerous substrates of CRL4L2DTL E3 ligase have been identified, such as CDT1, the CDK inhibitor, the histone methyltransferase Set8, the histone acetyltransferases GCN5, the checkpoint kinase CHK1 and the TOB anti-proliferative protein [54]. EED can strongly interact with CUL4A-DDB1 and participate in regulating the methylation of H3 histone at K9 and K27 [49]. However, EED protein sequence does not contain a recognizable WDxR/K motif, it has a WNIQ motif in the N- terminus of the WD40-repeat domain instead [49]. Deletion of any of the five putative WD40 repeats in EED abolishes its ability to mediate H3K27 methylation [55].
Additionally, the recent studies indicate that a WD40-repeat protein WDR18 can collaborate with DNA topoisomerase 2-binding protein 1 (TopBP1) to facilitate DNA damage checkpoint signaling [56]. WDR18 associates with the C-terminus of TopBP1 in vitro and in vivo, and the association between WDR18 and TopBP1 is required for AT70-induced Chk1 phosphorylation [56]. WDR18 facilitates ATR-dependent Chk1 phosphorylation via interacting with both C-terminus of TopBP1 and Chk1, promoting DNA damage checkpoint signaling [56].
3. WD40 proteins and cell death
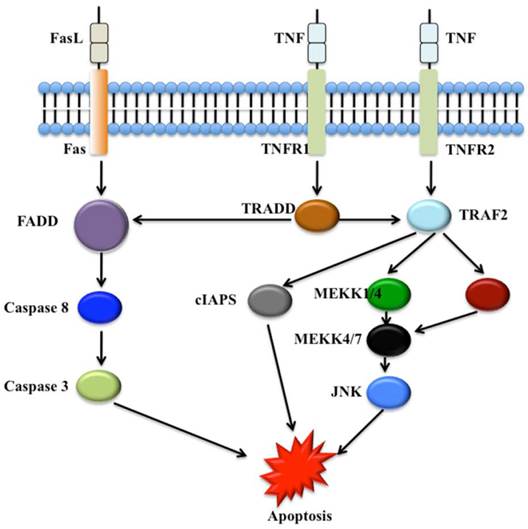
Numerous WD40 proteins have been revealed to involve cell death, in particular apoptosis and autophagy processes. In mammals, two main apoptotic initiation mechanisms have been suggested: the tumor necrosis factor (TNF) induced pathway and the Fas-Fas ligand-mediated pathway (Figure 3) [57]. Autophagy process involves cell degradation of unnecessary or dysfunctional cellular components through the actions of lysosomes [60, 61]. Three different forms of autophagy are commonly described: macroautophagy, microautophagy and chaperone-mediated autophagy [62].
The tumor necrosis factor (TNF) induced apoptosis pathway and the Fas-Fas ligand-mediated apoptosis pathway. The Fas, tumor necrosis factor receptor type 1 and 2 (TNFR1 and TNFR2) localize in the cell membrane. Numerous adaptor proteins and signal transducing molecules are involved in signal transduction initiated by the binding of the Fas ligand (FasL) to Fas and TNF to TNFR1 and TNFR2, respectively [58, 59]. cIAPs, cellular inhibitors of apoptosis; FADD, Fas-associated death domain protein; JNK, c-Jun N-terminal kinase; MEKK, Mitogen-activated protein kinase kinase kinase; TRADD, TNF receptor-associated death domain protein; TRAF2, TNF receptor-associated factor 2.
3.1 WD40 proteins and apoptosis
The intracellular caspases are required for the deliberate disassembly of cells into apoptotic bodies during apoptosis [63, 64]. Twelve caspases have been identified in human and they are present as inactive pro-enzymes that are activated by proteolytic cleavage [65]. Caspases3, 8, and 9 are function critically in apoptosis pathways [64]. Caspase3 can interact with both caspase8 and caspase9, and it appears to amplify caspase8 and caspase9 initiation signals into full-fledged commitment to disassembly [63]. Caspase8 and caspase9 activate caspase3 by proteolytic cleavage and caspase3 then cleaves vital cellular proteins or other caspases [66]. Caspase8 is responsible for initiating disassembly in response to extracellular apoptosis-inducing ligands [67, 68, 69]. Caspase9 activates disassembly in response to induction of stress signaling pathways including stress-activated protein kinase/c-Jun NH(2)-terminal kinase (SAPK/JNK) [70], which trigger the release of cytochrome c from mitochondria and activation of Apaf-1 [71]. Apaf-1 is a WD40 protein, and it is activated in the presence of dATP and cytochrome c binding, resulting in the activation of caspase9 [12]. The WD40 domain of Apaf-1 is located in the C-terminal region and plays critical role for its self-association and pro-caspase9 activation [72]. Interestingly, the WD40 repeats can interact with its N-terminal CED-4 homologous region [72]. Mutational analysis reveals that expression of the WD40 repeat region (residues 468-1194) inhibits Apaf-1 (residues 1-559) self-association and the maturation of pro-caspase9 [72]. The homologues of Apaf-1, including CED-4 in C. elegans and Dark in Drosophila, also can activate the caspases to initiate apoptosis [67, 73]. Interestingly, these proteins appear to have different regulation and biological roles [67]. Of them, Apaf-1 is inhibited by its WD40 repeats until it is activated by cytochrome c, derived from damaged mitochondria [67]. CED-4 is restrained by the homologue of B-cell lymphoma 2 (Bcl-2), namely, CED-9 [67]. The activation of Dark does not seem to involve cytochrome c, even though it also contains WD40 repeats [67].
WDR92 (also known as Monad) is a WD40 repeat protein and is widely expressed in human tissues [14]. Overexpression of WDR92 in HEK293 cells promotes apoptosis and caspase3 activation induced by TNF-α and cycloheximide, indicating that WDR92 may function as a novel modulator of apoptosis pathway [14]. WDR35 (also known as Naofen) is a novel WD40-repeat protein [13]. Overexpression of WDR35 in HEK293 cells spontaneously induces caspase3 activation and apoptosis, and exhibits extremely high susceptibility to TNF-α-induced apoptosis, suggesting that WDR35 may function as a novel modulator activating caspase3 and promote TNF-α-stimulated apoptosis [13]. However, there is no available information about the function of WD40 repeats in these two proteins.
The TRAF protein family members associate with, and mediate the signal transduction from members of TNF receptor superfamily [15]. Among these TRAF proteins, only TRAF7 contains seven WD40 repeats at its C-terminus [15]. In addition, TRAF7 also contains another important domain: RING finger [15]. The biochemical analysis indicates that its WD40 repeats specifically interacts with MEKK3 (MAP3K3) and induces MEKK3-mediated AP1 and CHOP activation [15]. The RING finger domain is required for its apoptotic activity as its deletion significantly reduces the ability to induce cell death [15].
Vps41 contains a WD40 domain, a CLH domain and a RING-finger motif mediating membrane association [17, 74]. Vps41 binds to caspase8 through its WD40 region and RING-finger motif [17]. Overexpression of Vps41 promotes Fas-induced apoptosis in A549 human lung adenocarcinoma cells [17].
The human immunodeficiency virus type 1 (HIV-1) Vpr-binding protein VprBP contains WD40 repeats [75]. VprBP has been found to mediate Vpr-induced G2 cell cycle arrest, and bind stoichiometrically with DDB1 through its WD40 domain to form a complex with CUL4A [75]. Conditional deletion of the VprBP gene in mouse embryonic fibroblasts results in severely defective progression through S phase and subsequent apoptosis [75].
LRRK2 also contains WD40 repeats [30], and mutations in LRRK2 cause autosomal-dominant Parkinson's disease (PD) [30]. The WD40 repeats are required for LRRK2 neurotoxicity and deletion of WD40 repeats completely blocks the neurotoxicity associated with LRRK2 PD mutations [76]. LRRK2 is also involved in cell death processes as it interacts with FADD (Fas-associated protein with dead domain), which is implicated in the activation of apoptosis [77]. Some studies have further indicated that FADD/caspase8 contributes to the apoptosis induced by LRRK2 [77].
The receptor for activated C kinase 1 (RACK1) is a member of WD40 proteins [78]. In response to stress, RACK1 is sequestered into stress granules and inhibits apoptosis by suppressing stress-responsive MAPK pathways [79]. The p53 family members such as p63 and p73 can induce cell cycle arrest and promote apoptosis through a transactivation (TA) domain [80]. ΔNp63 isotypes lacking the TA-domain promote cell proliferation and tumorigenesis in vitro and in vivo [80]. Interestingly, RACK1 physically associates with the p63α C-terminal domain through its WD40 domain, regulating apoptosis [80].
3.2 WD40 proteins and autophagy
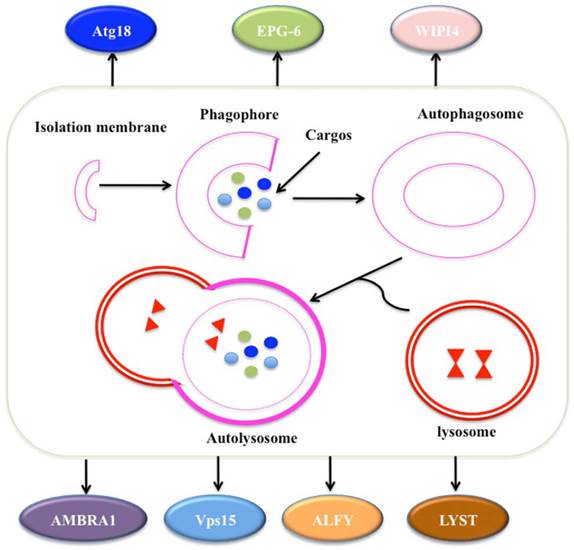
Autophagy is a self-degradative process involved in many aspects: (1) autophagy is initiated by the ULK1 (unc-51 like autophagy activating kinase 1) complex, which consists of ULK1, Atg13, Atg17 and receives stress signals from mTOR (mammalian target of rapamycin) complex 1 [81, 82]; (2) autophagosome formation is controlled by Beclin-1/Vps34 at membranes in response to stress signaling pathways [83, 84]; (3) Atg5-Atg12 conjugates and interacts with Atg16L, thereby polymerising at the autophagosome [83, 84]; (4) the LC3 (Microtubule-associated protein 1A/1B-light chain 3) system facilitates the transport and maturation of the autophagosome [85]; and (5) autophagosome fuses its external membrane with lysosomes to degrade its cargos (Figure 4) [86].
The process of autophagy. The process of macroautophagy consists of three main stages: initiation of the formation of the isolation membrane/phagophore; elongation and completion of the mature autophagosome; and fusion of the autophagosome with the lysosome to form an autolysosome [87]. A variety of WD40 proteins have been identified to involve this process, such as Atg18, EPG-6, WIPI4, AMBRA1, Vps15, ALFY and LYST.
Atg family proteins are required for autophagosome formation [88]. Among these Atg proteins, Atg18 protein contains a WD40 domain with 7 beta-propellers [89]. Phosphatidylinositol 3-phosphate (PtdIns(3)P), generated by the Vps34 PI(3)K complex, can recruit Atg18 to the preautophagosomal structure (PAS) in yeast and the mammalian Atg18 homologs, WIPI1 (WD repeat domain, phosphoinositide interacting 1) and WIPI2, to early autophagic structures [19, 90, 91, 92]. The WD40 domain in Atg18 contains a conserved FRRG motif, which binds to phosphoinositides and promotes Atg18 recruitment to the PAS [18]. In the absence of the FRRG motif, the inability of Atg18 interacting with Atg2 impairs its association with the PAS, causing an autophagy block [18]. The homolog of Atg18 in C. elegans, namely, EPG-6, is also a WD40 repeat protein with PtdIns(3)P-binding activity [19]. EPG-6 directly interacts with Atg2 through the sixth WD40 repeat, regulating progression of omegasomes to autophagosomes, and their loss of function causes accumulation of enlarged early autophagic structures [19]. The mammalian EPG-6 homolog, namely, WIPI4, is also essential for progression of omegasomes to autophagosomes in starvation-induced autophagy [19].
AMBRA1 is a WD40 protein that regulates autophagy and development of the nervous system [20]. AMBRA1 can interact with Atg6 and PIK3C3 (phosphatidylinositol 3-kinase, catalytic subunit type 3) [93, 94]. The decreased expression of AMBRA1 leads to autophagy impairment and associates with increased cell death [20]. The functional inactivation of AMBRA1 in mouse causes lethality in utero, leading to severe neural tube defects associated with autophagy impairment, accumulation of ubiquitinated proteins, and excessive apoptosis [20, 95]. The core Vps34/PI3K complex consists of Atg6, Vps15 and Vps34 [96]. This complex is critical for recruiting autophagy-related proteins to the preautophagosomal structure and for other vesicular trafficking processes [97]. Vps15 kinase contains WD40 repeats, and is required for both stress-induced and developmentally programmed autophagy in various tissues in Drosophila melanogaster [96]. The X-ray crystal structure of the Vps15 WD40 domain reveals a seven-bladed propeller resembling that of typical Gβ subunits [98]. The WD40 domain is sufficient to bind Gpa1 (Guanine nucleotide-binding protein alpha-1 subunit) and Atg14 [98].
A variety of autophagy proteins contain an LC3‐interacting region (LIR), which is required for their interaction with Atg8 homolog proteins [97]. ALFY (also known as WDFY3) binds selectively to LC3C and the GABARAPs (gamma-aminobutyric acid receptor-associated protein) through a LIR in its WD40 domain [97]. The ALFY‐GABARAP interaction is required for the clearance of certain p62‐bound aggregates by autophagy [97].
Most BEACH (named after 'Beige and Chediak-Higashi') domain-containing proteins (BDCPs) harbor WD40 repeats [99]. Studies indicate that mutations in individual BDCPs result in severe human diseases [99]. The human lysosomal trafficking regulator (LYST) is the most studied BDCP protein in human, which can affect lysosome size and quantity, but not trafficking or degradation through autophagy or endocytosis [100]. However, the homolog of LYST in Drosophila exhibits a critical role in the maturation of phagosomes and autophagosomes [101]. LRBA (Lipopolysaccharide-responsive and beige-like anchor protein) is also a WD40 repeat-containing BDCP protein. Mutations of LRBA cause an immune deficiency characterized by defects in B cell activation and autophagy and by susceptibility to apoptosis [102].
3.3 WD40 proteins and other types of cell death
Some other WD40 proteins are also involved in cell death, such as WDR26 and MDV1 (mitochondrial division, also known as Net2). Hydrogen peroxide (H2O2) can induce cell death and significantly up-regulate the expression of WDR26 [103], and WDR26 overexpression in turn strongly suppressed H2O2-induced cell death [103]. MDV1 is a multi-domain protein containing three major regions: an N-terminal unknown structure, a central coiled-coil (C-C) domain, and a C-terminal WD40 domain [104]. Studies indicate that the MDV1 interacts with Dnm1 (dynamin-related GTPase) through WD40 domain, and interact with Fis1 (Mitochondria fission 1 protein) via N-terminal region [104]. The MDV1-Dnm1-Fis1 complex has been demonstrated to regulate programmed cell death in the mitochondria in budding yeast Saccharomyces cerevisiae [104].
4. WD40 proteins and cell cycle control
Coordinated progression through the cell cycle is critical for eukaryotic cells [105]. Eukaryotic cell cycle progression is mainly controlled by a family of protein kinases known as cyclin-dependent kinases (CDKs), which consist of an activating cyclin subunit and a catalytic subunit CDK [39, 106]. The CDKs activities are controlled by the availability of their cyclin partners and the expression of specific CDK inhibitors (CDKIs) [103]. Interestingly, WD40 proteins have emerged as crucial regulators of cell cycle progression via interacting with other proteins.
Cell cycle progression is tightly controlled by UPS, as cyclin expression levels are executed via UPS and CDKIs are targeted for degradation by the UPS-mediated ubiquitination [107]. Two ubiquitin ligases, namely, SKP1-CUL1-F-box-protein (SCF) complex and anaphase-promoting complex/cyclosome (APC/C), specifically function in the ubiquitination of many of these CDKs and CDKIs [107]. SCF can control the transition of G1/S, and two key targets of SCF, cyclin E and the CDKI p27Kip1, are controlled by CUL4-DDB1 [49, 108]. In addition, several F-box proteins with WD40 repeats, such as FBXW1 (also known as β-TRCP), FBXW5, FBXW7 and FBXW8, can bind DDB1 through WD40 repeats in vitro in respond to DNA damage, repair and cell cycle progression [22]. FBXW1 plays important roles in regulating cell cycle checkpoints [109]. In response to genotoxic stress, it contributes to turn off CDK1 activity by mediating the degradation of CDC25A in collaboration with Chk1 [109, 110], thereby preventing cell cycle progression before the completion of DNA repair [109, 110]. FBXW5 is a cell cycle-regulated protein with expression levels peaking at the G1/S transition [111]. FBXW5 levels are controlled by the anaphase-promoting (APC/C) complex, which targets FBXW5 for degradation during mitosis and G1, thereby helping to reset the centrosome duplication machinery [111]. FBXW7 protein is a well-established tumor suppressor and responsible for substrate recognition in an SCF ubiquitin ligase complex [24, 28]. The WD40 repeats in FBXW7 is essential for binding substrates such as cyclin E, Myc (myelocytomatosis oncogene), JUN (jun proto-oncogene) and Notch [24]. FBXW8 plays an essential role in cancer cell proliferation through proteolysis of cyclin D1, but it remains unclear whether FBXW8 is necessary for cell cycle progression in normal cells [25]. The CRL4AL2DTL E3 ligase also interacts with and targets the tumor suppressor protein p53, mediating cell cycle arrest or apoptosis in response to genotoxic stress [112].
CDC20 is a WD40 protein, which is an essential cell-cycle regulator required for the completion of mitosis in organisms from yeast to mammals [26]. In mitosis, CDC20 binds to and activates the ubiquitin ligase activity of APC/C and enables the ubiquitination and degradation of securin and cyclin B [20]. APC/CCDC20 contributes to the proteolysis of securin, triggering chromosomal separation at anaphase [107]. The structure of WD40 domain in CDC20 indicates that it folds into a seven-bladed β-propeller and has an ideal architecture for multiple protein-protein interactions [113]. Thus, CDC20 appears to bridge the interactions between APC/C and its substrates through WD40 repeats [113].
In budding yeast Saccharomyces cerevisiae, Sic1 encodes a cyclin-dependent kinase inhibitor that regulates the cell cycle at the G1 to S transition by inhibiting the activity of the CDC28 [114]. The phosphorylated Sic1 binds to a WD40 protein, namely, CDC4 (also known as FBXW7 in mammals), which is the substrate recognition subunit of the SCFCDC4 E3 ligase [115]. Versions of Sic1 that lack multiple CDK phosphorylation sites fail to bind CDC4, are stabilized in vivo, and cause cell cycle arrest [116, 117]. Sic1 binds to the WD40 domain of CDC4 when a nominal set of six of the nine CDK consensus sites is phosphorylated, terming CDC4 phosphodegron (CPDs) [118]. The human CDC4 ortholog FBXW7 shares similar phosphodegron to that of CDC4, with the exception of an additional phosphorylated residue at the P + 4 position [119, 120].
5. Conclusion
Genome integrity and cell cycle progression are tightly regulated by a variety of pathways that mainly involve DNA repair and recombination, apoptosis, autophagy and cell cycle checkpoints. Numerous WD40 proteins participate in these processes through the following mechanisms: (1) function as substrate receptors of CRL4 E3 ligases to target different substrates selectively [1]; (2) utilization of insertion motifs and inter-blade binding grooves of WD40 domains in ligand binding [1]; and (3) recognition of post-translational modification marks [1]. Considering the critical roles of WD40 proteins in genome stability and cell cycle progression, it is clear that there is much more work to be investigated about the E3 ligase substrates and WD40 interaction proteins specifically responding to DNA damage and repair, apoptosis, autophagy and cell cycle progression in the future.
Acknowledgements
We apologize to all authors whose contributions were not cited due to space limitations. We would like to express our gratitude to Dr. Jianbin Wang for critically reading the manuscript and for facilitating discussions.
Compliance with Ethics Guidelines
This article does not contain any studies with human or animal as subjects performed by the author.
Competing Interests
The authors have declared that no competing interest exists.
References
1. Xu C, Min J. Structure and function of WD40 domain proteins. Protein Cell. 2011;2:202-14
2. Letunic I, Doerks T, Bork P. SMART: recent updates, new developments and status in 2015. Nucleic Acids Res. 2014 doi: 10.1093/nar/gku949
3. Ouyang Y, Huang X, Lu Z, Yao J. Genomic survey, expression profile and co-expression network analysis of OsWD40 family in rice. BMC Genomics. 2012;13:100
4. van Nocker S, Ludwig P. The WD-repeat protein superfamily in Arabidopsis: conservation and divergence in structure and function. BMC Genomics. 2003;4:50
5. Suganuma T, Pattenden SG, Workman JL. Diverse functions of WD40 repeat proteins in histone recognition. Genes Dev. 2008;22:1265-8
6. Mishra AK, Puranik S, Bahadur RP, Prasad M. The DNA-binding activity of an AP2 protein is involved in transcriptional regulation of a stress-responsive gene, SiWD40, in foxtail millet. Genomics. 2012;100:252-63
7. Jin J, Arias EE, Chen J. et al. A family of diverse Cul4-Ddb1-interacting proteins includes Cdt2, which is required for S phase destruction of the replication factor Cdt1. Mol Cell. 2006;23:709-21
8. Zhang C, Guo H, Zhang J. et al. Arabidopsis cockayne syndrome A-like proteins 1A and 1B form a complex with CULLIN4 and damage DNA binding protein 1A and regulate the response to UV irradiation. Plant Cell. 2010;22:2353-69
9. Lee J, Zhou P. DCAFs, the missing link of the CUL4-DDB1 ubiquitin ligase. Mol Cell. 2007;26:775-80
10. Havens CG, Walter JC. Mechanism of CRL4(Cdt2), a PCNA-dependent E3 ubiquitin ligase. Genes & development. 2011;25:1568-82
11. Higa LA, Wu M, Ye T. et al. CUL4-DDB1 ubiquitin ligase interacts with multiple WD40-repeat proteins and regulates histone methylation. Nat Cell Biol. 2006;8:1277-83
12. Li P, Nijhawan D, Budihardjo I. et al. Cytochrome c and dATP-dependent formation of Apaf-1/caspase-9 complex initiates an apoptotic protease cascade. Cell. 1997;91:479-89
13. Feng GG, Li C, Huang L. et al. Naofen, a novel WD40-repeat protein, mediates spontaneous and tumor necrosis factor-induced apoptosis. Biochem Biophys Res Commun. 2010;394:153-7
14. Saeki M, Irie Y, Ni L. et al. Monad, a WD40 repeat protein, promotes apoptosis induced by TNF-alpha. Biochem Biophys Res Commun. 2006;342:568-72
15. Xu LG, Li LY, Shu HB. TRAF7 potentiates MEKK3-induced AP1 and CHOP activation and induces apoptosis. J Biol Chem. 2004;279:17278-82
16. Ho CC, Rideout HJ, Ribe E. et al. The Parkinson disease protein leucine-rich repeat kinase 2 transduces death signals via Fas-associated protein with death domain and caspase-8 in a cellular model of neurodegeneration. J Neurosci. 2009;29:1011-6
17. Wang L, Pan X, He L. et al. Vps41, a protein involved in lysosomal trafficking, interacts with caspase-8. Acta Biochim Pol. 2013;60:37-42
18. Rieter E, Vinke F, Bakula D. et al. Atg18 function in autophagy is regulated by specific sites within its beta-propeller. J Cell Sci. 2013;126:593-604
19. Lu Q, Yang P, Huang X. et al. The WD40 repeat PtdIns(3)P-binding protein EPG-6 regulates progression of omegasomes to autophagosomes. Dev Cell. 2011;21:343-57
20. Fimia GM, Stoykova A, Romagnoli A. et al. Ambra1 regulates autophagy and development of the nervous system. Nature. 2007;447:1121-5
21. Lystad AH, Ichimura Y, Takagi K. et al. Structural determinants in GABARAP required for the selective binding and recruitment of ALFY to LC3B-positive structures. EMBO Rep. 2014;15:557-65
22. He YJ, McCall CM, Hu J. et al. DDB1 functions as a linker to recruit receptor WD40 proteins to CUL4-ROC1 ubiquitin ligases. Genes Dev. 2006;20:2949-54
23. Puklowski A, Homsi Y, Keller D. et al. The SCF-FBXW5 E3-ubiquitin ligase is regulated by PLK4 and targets HsSAS-6 to control centrosome duplication. Nat Cell Biol. 2011;13:1004-9
24. Takeishi S, Nakayama KI. Role of Fbxw7 in the maintenance of normal stem cells and cancer-initiating cells. Br J Cancer. 2014;111:1054-9
25. Okabe H, Lee SH, Phuchareon J. et al. A critical role for FBXW8 and MAPK in cyclin D1 degradation and cancer cell proliferation. PLoS One. 2006;1:e128
26. Yu H. Cdc20: a WD40 activator for a cell cycle degradation machine. Mol Cell. 2007;27:3-16
27. Henning KA, Li L, Iyer N. et al. The Cockayne syndrome group A gene encodes a WD repeat protein that interacts with CSB protein and a subunit of RNA polymerase II TFIIH. Cell. 1995;82:555-64
28. Wang Z, Liu P, Inuzuka H, Wei W. Roles of F-box proteins in cancer. Nat Rev Cancer. 2014;14:233-47
29. Vallee RB, Tsai JW. The cellular roles of the lissencephaly gene LIS1, and what they tell us about brain development. Genes Dev. 2006;20:1384-93
30. Zimprich A, Biskup S, Leitner P. et al. Mutations in LRRK2 cause autosomal-dominant parkinsonism with pleomorphic pathology. Neuron. 2004;44:601-7
31. Jackson SP, Durocher D. Regulation of DNA damage responses by ubiquitin and SUMO. Mol Cell. 2013;49:795-807
32. Rastogi RP. et al. Molecular mechanisms of ultraviolet radiation-induced DNA damage and repair. J Nucleic Acids. 2010;2010:592980
33. Davis AJ, Chen DJ. DNA double strand break repair via non-homologous end-joining. Transl Cancer Res. 2013;2:130-43
34. D'Errico M, Teson M, Calcagnile A. et al. Apoptosis and efficient repair of DNA damage protect human keratinocytes against UVB. Cell Death Differ. 2003;10:754-56
35. Sato K, Sundaramoorthy E, Rajendra E. et al. A DNA-damage selective role for BRCA1 E3 ligase in claspin ubiquitylation, CHK1 activation, and DNA repair. Curr Biol. 2012;22:1659-66
36. Lecker SH, Goldberg AL, Mitch WE. Protein degradation by the ubiquitin-proteasome pathway in normal and disease states. J Am Soc Nephrol. 2006;17:1807-19
37. Wei D, Sun Y. Small RING Finger Proteins RBX1 and RBX2 of SCF E3 Ubiquitin Ligases: The Role in Cancer and as Cancer Targets. Genes & cancer. 2010:700-7
38. Zhao Y, Sun Y. Cullin-RING Ligases as attractive anti-cancer targets. Current pharmaceutical design. 2013;19:3215-25
39. Higa LA, Zhang H. Stealing the spotlight: CUL4-DDB1 ubiquitin ligase docks WD40-repeat proteins to destroy. Cell Div. 2007;2:5
40. Bertola DR, Cao H, Albano LM. et al. Cockayne syndrome type A: novel mutations in eight typical patients. J Hum Genet. 2006;51:701-5
41. Hoeijmakers JH. DNA damage, aging, and cancer. N Engl J Med. 2009;361:1475-85
42. Laugel V, Dalloz C, Durand M. et al. Mutation update for the CSB/ERCC6 and CSA/ERCC8 genes involved in Cockayne syndrome. Hum Mutat. 2010;31:113-26
43. Groisman R, Kuraoka I, Chevallier O. et al. CSA-dependent degradation of CSB by the ubiquitin-proteasome pathway establishes a link between complementation factors of the Cockayne syndrome. Genes & development. 2006;20:1429-34
44. Scrima A, Konickova R, Czyzewski BK. et al. Structural basis of UV DNA-damage recognition by the DDB1-DDB2 complex. Cell. 2008;135:1213-23
45. Hannah J, Zhou P. Regulation of DNA damage response pathways by the cullin-RING ubiquitin ligases. DNA Repair (Amst). 2009;8:536-43
46. Fei J, Kaczmarek N, Luch A. et al. Regulation of nucleotide excision repair by UV-DDB: prioritization of damage recognition to internucleosomal DNA. PLoS Biol. 2011;9:e1001183
47. Fischer ES, Scrima A, Bohm K. et al. The molecular basis of CRL4DDB2/CSA ubiquitin ligase architecture, targeting, and activation. Cell. 2011;147:1024-39
48. Zhang L, Lubin A, Chen H. et al. The deubiquitinating protein USP24 interacts with DDB2 and regulates DDB2 stability. Cell cycle. 2012;11:4378-84
49. Higa LA, Yang X, Zheng J. et al. Involvement of CUL4 ubiquitin E3 ligases in regulating CDK inhibitors Dacapo/p27Kip1 and cyclin E degradation. Cell Cycle. 2006;5:71-7
50. Trievel RC, Shilatifard A. WDR5, a complexed protein. Nature structural & molecular biology. 2009;16:678-80
51. Foucher I, Montesinos ML, Volovitch M. et al. Joint regulation of the MAP1B promoter by HNF3beta/Foxa2 and Engrailed is the result of a highly conserved mechanism for direct interaction of homeoproteins and Fox transcription factors. Development. 2003;130:1867-76
52. Anastas JN, Moon RT. WNT signalling pathways as therapeutic targets in cancer. Nature reviews Cancer. 2013;13:11-26
53. Daniels DL, Weis WI. Beta-catenin directly displaces Groucho/TLE repressors from Tcf/Lef in Wnt-mediated transcription activation. Nature structural & molecular biology. 2005;12:364-371
54. Olivero M, Dettori D, Arena S. et al. The stress phenotype makes cancer cells addicted to CDT2, a substrate receptor of the CRL4 ubiquitin ligase. Oncotarget. 2014;5:5992-6002
55. Montgomery ND, Yee D, Montgomery SA, Magnuson T. Molecular and functional mapping of EED motifs required for PRC2-dependent histone methylation. Journal of molecular biology. 2007;374:1145-57
56. Yan S, Willis J. WD40-repeat protein WDR18 collaborates with TopBP1 to facilitate DNA damage checkpoint signaling. Biochem Biophys Res Commun. 2013;431:466-71
57. Waring P, Mullbacher A. Cell death induced by the Fas/Fas ligand pathway and its role in pathology. Immunol Cell Biol. 1999;77:312-7
58. Mak TW, Yeh WC. Signaling for survival and apoptosis in the immune system. Arthritis Res. 2002;4:243-52
59. Wu Y, Zhou BP. TNF-alpha/NF-kappaB/Snail pathway in cancer cell migration and invasion. Br J Cancer. 2010;102:639-44
60. Kalamida D, Karagounis IV, Giatromanolaki A, Koukourakis MI. Important role of autophagy in endothelial cell response to ionizing radiation. PLoS One. 2014;9:e102408
61. Papackova Z, Cahova M. Important role of autophagy in regulation of metabolic processes in health, disease and aging. Physiol Res. 2014;63:409-20
62. Kaushik S, Massey AC, Mizushima N, Cuervo AM. Constitutive activation of chaperone-mediated autophagy in cells with impaired macroautophagy. Mol Biol Cell. 2008;19:2179-92
63. Mallat Z, Tedgui A. Apoptosis in the vasculature: mechanisms and functional importance. Br J Pharmacol. 2000;130:947-62
64. Yu FS, Yu CS, Chen JC. et al. Tetrandrine induces apoptosis Via caspase-8, -9, and -3 and poly (ADP ribose) polymerase dependent pathways and autophagy through beclin-1/ LC3-I, II signaling pathways in human oral cancer HSC-3 cells. Environ Toxicol. 2014 doi:10.1002/tox.22053
65. McIlwain DR, Berger T, Mak TW. Caspase functions in cell death and disease. Cold Spring Harb Perspect Biol. 2013;5:a008656
66. Parrish AB, Freel CD, Kornbluth S. Cellular mechanisms controlling caspase activation and function. Cold Spring Harb Perspect Biol. 2013 5
67. Adams JM, Cory S. Apoptosomes: engines for caspase activation. Curr Opin Cell Biol. 2002;14:715-20
68. Elmore S. Apoptosis: a review of programmed cell death. Toxicol Pathol. 2007;35:495-516
69. Ouyang L, Shi Z, Zhao S. et al. Programmed cell death pathways in cancer: a review of apoptosis, autophagy and programmed necrosis. Cell Prolif. 2012;45:487-98
70. Panaretakis T, Laane E, Pokrovskaja K. et al. Doxorubicin requires the sequential activation of caspase-2, protein kinase Cdelta, and c-Jun NH2-terminal kinase to induce apoptosis. Mol Biol Cell. 2005;16:3821-31
71. Martinou JC, Desagher S, Antonsson B. Cytochrome c release from mitochondria: all or nothing. Nat Cell Biol. 2000;2:E41-3
72. Hu Y, Ding L, Spencer DM, Nunez G. WD-40 repeat region regulates Apaf-1 self-association and procaspase-9 activation. J Biol Chem. 1998;273:33489-94
73. Kornbluth S, White K. Apoptosis in Drosophila: neither fish nor fowl (nor man, nor worm). J Cell Sci. 2005;118:1779-87
74. Ruan Q, Harrington AJ, Caldwell KA. et al. VPS41, a protein involved in lysosomal trafficking, is protective in Caenorhabditis elegans and mammalian cellular models of Parkinson's disease. Neurobiol Dis. 2010;37:330-8
75. McCall CM, Miliani de Marval PL. et al. Human immunodeficiency virus type 1 Vpr-binding protein VprBP, a WD40 protein associated with the DDB1-CUL4 E3 ubiquitin ligase, is essential for DNA replication and embryonic development. Mol Cell Biol. 2008;28:5621-33
76. Jorgensen ND, Peng Y, Ho CC. et al. The WD40 domain is required for LRRK2 neurotoxicity. PLoS One. 2009;4:e8463
77. Ho CC, Rideout HJ, Ribe E. et al. The Parkinson disease protein leucine-rich repeat kinase 2 transduces death signals via Fas-associated protein with death domain and caspase-8 in a cellular model of neurodegeneration. J Neurosci. 2009;29:1011-6
78. Adams DR, Ron D, Kiely PA. RACK1, A multifaceted scaffolding protein: Structure and function. Cell Commun Signal. 2011;9:22
79. Arimoto K, Fukuda H, Imajoh-Ohmi S. et al. Formation of stress granules inhibits apoptosis by suppressing stress-responsive MAPK pathways. Nat Cell Biol. 2008;10:1324-32
80. Fomenkov A, Zangen R, Huang YP. et al. RACK1 and stratifin target DeltaNp63alpha for a proteasome degradation in head and neck squamous cell carcinoma cells upon DNA damage. Cell Cycle. 2004;3:1285-95
81. Jung CH, Jun CB, Ro SH. et al. ULK-Atg13-FIP200 complexes mediate mTOR signaling to the autophagy machinery. Mol Biol Cell. 2009;20:1992-2003
82. Mizushima N. The role of the Atg1/ULK1 complex in autophagy regulation. Curr Opin Cell Biol. 2010;22:132-9
83. Fimia GM, Stoykova A, Romagnoli A. et al. Ambra1 regulates autophagy and development of the nervous system. Nature. 2007;447:1121-5
84. Glick D, Barth S, Macleod KF. Autophagy: cellular and molecular mechanisms. J Pathol. 2010;221:3-12
85. Matsushita M, Suzuki NN, Obara K. et al. Structure of Atg5.Atg16, a complex essential for autophagy. J Biol Chem. 2007;282:6763-72
86. Pattingre S, Espert L, Biard-Piechaczyk M, Codogno P. Regulation of macroautophagy by mTOR and Beclin 1 complexes. Biochimie. 2008;90:313-23
87. Vellai T. Autophagy genes and ageing. Cell Death Differ. 2009;16:94-102
88. Nakatogawa H, Suzuki K, Kamada Y, Ohsumi Y. Dynamics and diversity in autophagy mechanisms: lessons from yeast. Nat Rev Mol Cell Biol. 2009;10:458-67
89. Dove SK, Piper RC, McEwen RK. et al. Svp1p defines a family of phosphatidylinositol 3,5-bisphosphate effectors. EMBO J. 2004;23:1922-33
90. Obara K, Sekito T, Niimi K, Ohsumi Y. The Atg18-Atg2 complex is recruited to autophagic membranes via phosphatidylinositol 3-phosphate and exerts an essential function. J Biol Chem. 2008;283:23972-80
91. Proikas-Cezanne T, Ruckerbauer S. et al. Human WIPI-1 puncta-formation: a novel assay to assess mammalian autophagy. FEBS Lett. 2007;581:3396-404
92. Polson HE, de Lartigue J, Rigden DJ. et al. Mammalian Atg18 (WIPI2) localizes to omegasome-anchored phagophores and positively regulates LC3 lipidation. Autophagy. 2010;6:506-22
93. Cao Y, Klionsky DJ. Physiological functions of Atg6/Beclin 1: a unique autophagy-related protein. Cell Res. 2007;17:839-49
94. Xia P, Wang S, Huang G. et al. RNF2 is recruited by WASH to ubiquitinate AMBRA1 leading to downregulation of autophagy. Cell Res. 2014;24:943-58
95. Cecconi F, Di Bartolomeo S, Nardacci R. et al. A novel role for autophagy in neurodevelopment. Autophagy. 2007;3:506-8
96. Anding AL, Baehrecke EH. Vps15 is required for stress induced and developmentally triggered autophagy and salivary gland protein secretion in Drosophila. Cell Death Differ. 2014 doi:10.1038/cdd.2014.174
97. Lystad AH, Ichimura Y, Takagi K. et al. Structural determinants in GABARAP required for the selective binding and recruitment of ALFY to LC3B-positive structures. EMBO Rep. 2014;15:557-65
98. Heenan EJ, Vanhooke JL, Temple BR. et al. Structure and function of Vps15 in the endosomal G protein signaling pathway. Biochemistry. 2009;48:6390-401
99. Cullinane AR, Schaffer AA, Huizing M. The BEACH is hot: a LYST of emerging roles for BEACH-domain containing proteins in human disease. Traffic. 2013;14:749-66
100. Holland P, Torgersen ML, Sandvig K, Simonsen A. LYST Affects Lysosome Size and Quantity, but not Trafficking or Degradation Through Autophagy or Endocytosis. Traffic. 2014 doi:10.1111/tra.12227
101. Rahman M, Haberman A, Tracy C. et al. Drosophila mauve mutants reveal a role of LYST homologs late in the maturation of phagosomes and autophagosomes. Traffic. 2012;13:1680-92
102. Lopez-Herrera G, Tampella G, Pan-Hammarstrom Q. et al. Deleterious mutations in LRBA are associated with a syndrome of immune deficiency and autoimmunity. Am J Hum Genet. 2012;90:986-1001
103. Zhao J, Liu Y, Wei X. et al. A novel WD-40 repeat protein WDR26 suppresses H2O2-induced cell death in neural cells. Neurosci Lett. 2009;460:66-71
104. Fannjiang Y, Cheng WC, Lee SJ. et al. Mitochondrial fission proteins regulate programmed cell death in yeast. Genes Dev. 2004;18:2785-97
105. Reinhardt HC, Yaffe MB. Phospho-Ser/Thr-binding domains: navigating the cell cycle and DNA damage response. Nat Rev Mol Cell Biol. 2013;14:563-80
106. Zhang C. Essential functions of iron-requiring proteins in DNA replication, repair and cell cycle control. Protein & cell. 2014 doi:10.1007/s13238-014-0083-7
107. Nakayama KI, Nakayama K. Ubiquitin ligases: cell-cycle control and cancer. Nat Rev Cancer. 2006;6:369-81
108. Sherr CJ, Roberts JM. CDK inhibitors: positive and negative regulators of G1-phase progression. Genes Dev. 1999;13:1501-12
109. Busino L, Donzelli M, Chiesa M. et al. Degradation of Cdc25A by beta-TrCP during S phase and in response to DNA damage. Nature. 2003;426:87-91
110. Jin J, Shirogane T, Xu L. et al. SCFbeta-TRCP links Chk1 signaling to degradation of the Cdc25A protein phosphatase. Genes Dev. 2003;17:3062-74
111. Puklowski A, Homsi Y, Keller D. et al. The SCF-FBXW5 E3-ubiquitin ligase is regulated by PLK4 and targets HsSAS-6 to control centrosome duplication. Nat Cell Biol. 2011;13:1004-9
112. Brooks CL, Gu W. p53 ubiquitination: Mdm2 and beyond. Mol Cell. 2006;21:307-15
113. Tian W, Li B, Warrington R. et al. Structural analysis of human Cdc20 supports multisite degron recognition by APC/C. Proc Natl Acad Sci U S A. 2012;109:18419-124
114. Schwob E, Bohm T, Mendenhall MD, Nasmyth K. The B-type cyclin kinase inhibitor p40SIC1 controls the G1 to S transition in S. cerevisiae. Cell. 1994;79:233-44
115. Wysocki R, Javaheri A, Kristjansdottir K. et al. CDK Pho85 targets CDK inhibitor Sic1 to relieve yeast G1 checkpoint arrest after DNA damage. Nat Struct Mol Biol. 2006;13:908-14
116. Tang X, Orlicky S, Mittag T. et al. Composite low affinity interactions dictate recognition of the cyclin-dependent kinase inhibitor Sic1 by the SCFCdc4 ubiquitin ligase. Proc Natl Acad Sci U S A. 2012;109:3287-92
117. Verma R, Annan RS, Huddleston MJ. et al. Phosphorylation of Sic1p by G1 Cdk required for its degradation and entry into S phase. Science. 1997;278:455-60
118. Nash P, Tang X, Orlicky S. et al. Multisite phosphorylation of a CDK inhibitor sets a threshold for the onset of DNA replication. Nature. 2001;414:514-21
119. Welcker M, Singer J, Loeb KR. et al. Multisite phosphorylation by Cdk2 and GSK3 controls cyclin E degradation. Mol Cell. 2003;12:381-92
120. Welcker M, Clurman BE. FBW7 ubiquitin ligase: a tumour suppressor at the crossroads of cell division, growth and differentiation. Nat Rev Cancer. 2008;8:83-93
121. Han Z, Xing X, Hu M. et al. Structural basis of EZH2 recognition by EED. Structure. 2007;15:1306-15
122. Pickles LM, Roe SM, Hemingway EJ. et al. Crystal structure of the C-terminal WD40 repeat domain of the human Groucho/TLE1 transcriptional corepressor. Structure. 2002;10:751-61
123. Reubold TF, Wohlgemuth S, Eschenburg S. Crystal structure of full-length Apaf-1: how the death signal is relayed in the mitochondrial pathway of apoptosis. Structure. 2011;19:1074-83
124. Mill P, Lockhart PJ, Fitzpatrick E. et al. Human and mouse mutations in WDR35 cause short-rib polydactyly syndromes due to abnormal ciliogenesis. Am J Hum Genet. 2011;88:508-15
125. Schwefel D, Groom HC, Boucherit VC. et al. Structural basis of lentiviral subversion of a cellular protein degradation pathway. Nature. 2014;505:234-38
126. Watanabe Y, Kobayashi T, Yamamoto H. et al. Structure-based analyses reveal distinct binding sites for Atg2 and phosphoinositides in Atg18. J Biol Chem. 2012;287:31681-90
127. Wu G, Xu G, Schulman BA. et al. Structure of a beta-TrCP1-Skp1-beta-catenin complex: destruction motif binding and lysine specificity of the SCF(beta-TrCP1) ubiquitin ligase. Mol Cell. 2003;11:1445-56
128. Puklowski A, Homsi Y, Keller D. et al. The SCF-FBXW5 E3-ubiquitin ligase is regulated by PLK4 and targets HsSAS-6 to control centrosome duplication. Nat Cell Biol. 2011;13:1004-9
129. Hao B, Oehlmann S, Sowa ME. et al. Structure of a Fbw7-Skp1-cyclin E complex: multisite-phosphorylated substrate recognition by SCF ubiquitin ligases. Mol Cell. 2007;26:131-43
Author contact
Corresponding author: caiguo.zhangedu

 Global reach, higher impact
Global reach, higher impact