ISSN: 1839-9940
J Genomics 2015; 3:59-71. doi:10.7150/jgen.10924 This volume Cite
Research Paper
Map-Based Comparative Genomic Analysis of Virulent Haemophilus Parasuis Serovars 4 and 5
Newport Laboratories Inc., Worthington, Minnesota, United States of America.
Published 2015-4-1
Abstract
Haemophilus parasuis is a commensal bacterium of the upper respiratory tract of healthy pigs. However, in conjunction with viral infections in immunocompromised animals H. parasuis can transform into a pathogen that is responsible for causing Glasser's disease which is typically characterized by fibrinous polyserositis, polyarthritis, meningitis and sometimes acute pneumonia and septicemia in pigs. Haemophilus parasuis serovar 5 is highly virulent and more frequently isolated from respiratory and systemic infection in pigs. Recently a highly virulent H. parasuis serovar 4 was isolated from the tissues of diseased pigs. To understand the differences in virulence and virulence-associated genes between H. parasuis serovar 5 and highly virulent H. parasuis serovar 4 strains, a genomic library was generated by TruSeq preparation and sequenced on Illumina HiSeq 2000 obtaining 50 bp PE reads. A three-way comparative genomic analysis was conducted between two highly virulent H. parasuis serovar 4 strains and H. parasuis serovar 5. Haemophilus parasuis serovar 5 GenBank isolate SH0165 (GenBank accession number CP001321.1) was used as reference strain for assembly.
Results of these analysis revealed the highly virulent H. parasuis serovar 4 lacks genes encoding for, glycosyl transferases, polysaccharide biosynthesis protein capD, spore coat polysaccharide biosynthesis protein C, polysaccharide export protein and sialyltransferase which can modify the lipopolysaccharide forming a short-chain LPS lacking O-specific polysaccharide chains often referred to as lipooligosaccharide (LOS). In addition, it can modify the outer membrane protein (OMP) structure. The lack of sialyltransferase significantly reduced the amount of sialic acid incorporated into LOS, a major and essential component of the cell wall and an important virulence determinant. These molecules may be involved in various stages of pathogenesis through molecular mimicry and by causing host cell cytotoxicity, reduced inflammatory and immunological response to infection with this organism. The mechanism by which sialyation of LPS contributes to virulence is a key to understanding the pathogenesis of this highly virulent H. parasuis serovar 4. This analysis also revealed the presence of virulence associated genes similar to the MerR family transcriptional regulators, macrophage infectivity potentiator protein, hemolysin, opacity associated protein, toxin antitoxin system, and virulence associated protein D and colicins. Haemophilus parasuis serovar 4 variants also possess extensive metal ion uptake and regulation mechanism which controls various virulence and virulence associated genes. A combination of virulence associated factors and/or genes and proteins with overlapping functions may be responsible for the apparent enhanced virulence of this organism.
The extensive structural modification of LOS and OMP of variant H. parasuis serovar 4 strains appear to aid in nasal colonization, are associated with the organisms' ability to evade the host immune response and provide serum-resistance. In addition, the combination of capsule modification and phase variation due to LOS substitutions could help variant H. parasuis serovar 4 transform into a highly virulent pathogen. Based on these results, the variant H. parasuis serovar 4 strains harbor a diverse repertoire of virulence associated genes which have not been previously reported.
Keywords: Haemophilus parasuis, sialic acid, sialyltransferase, lipooligosaccharide, outer membrane proteins.
Introduction
Haemophilus parasuis is a γ-proteobacteria, which belongs to the Pasteurellaceae family, is a rod-shaped, Gram-negative, non-motile, non-haemolytic, pleomorphic and nicotinamide adenine dinucleotide (NAD)-dependent bacterium (1, 2). It is a commensal bacterium of the upper respiratory tract of healthy pigs (3, 4). In conjunction with viral infections in immunocompromised animals, H. parasuis can transform into a pathogen responsible for causing Glasser's disease, which is typically characterized by fibrinous polyserositis, polyarthritis, meningitis and sometimes acute pneumonia and septicemia (3-5). However, clinical signs of H. parasuis infection in diseased pigs are mostly non-specific. This disease is one of the primary causes of morbidity and mortality in the U.S. and swine industries worldwide, resulting in substantial economic losses.
To date, 15 different serovars of H. parasuis have been identified based on the presence of heat stable antigens and gel diffusion tests, although a high percentage of the field isolates are non-typable (6). However, the highly discriminatory multilocus variable number of tandem repeats analysis (MLVA) may reduce the number of non-typable isolates (7). Different serovars of H. parsuis exhibit different degrees of virulence, ranging from highly virulent to non-virulent. Of the 15 recognized serovars, H. parasuis serovar 5 is more frequently isolated from respiratory and systemic infection in pigs (8). In addition, H. parasuis serovar 5 is one of the major causative agents of neonatal mortality in the pig industry worldwide (9). During the past few years, Newport Laboratories has observed an increase in the number of cases of H. parasuis serovar 4 infections in the U.S. Recent surveys have also reported H. parasuis serovar 4 is an emerging pathogen in the U.S. with a potential to cause serious disease outbreaks (10).
Like other members of Pasteurellaceae, H. parasuis has a large number of virulence and virulence-associated genes. These include lipopolysaccharide (LPS), capsular polysaccharide, adhesins/fimbriae, outer membrane proteins, neuraminidase, iron and heavy metal acquisition and transport systems (3, 11, 12, 13, 14). However, there is no direct correlation between the expression of these virulence factors and the virulence of H. parasuis isolates. In addition, the molecular basis underlying the expression of candidate virulence factors is not yet fully elucidated due to the complex gene regulatory mechanisms. Various serovars of H. parasuis exhibit a high heterogeneity at the molecular level, mostly due to recombination or lateral gene transfer.
We have isolated several highly virulent H. parasuis serovar 4 strains from the lungs of diseased pigs, which died of severe pneumonia and septicemia. Highly virulent H. parasuis strains can be opportunistic pathogens secondary to pneumonia, viral infection and/or stress causing septicemia without polyserositis or Glässer's disease which is characterized by polyserositis, pericarditis, arthritis and meningitis (3,4,5). To understand the differences in virulence and virulence-associated genes between H. parasuis serovar 5 and highly virulent H. parasuis serovars 4 strains we performed a three-way comparative genomic analysis between two H. parasuis serovar 4 strains and a virulent H. parasuis serovar 5 a U.S. isolate.
Results and discussion
Gene coverage and overall mapping
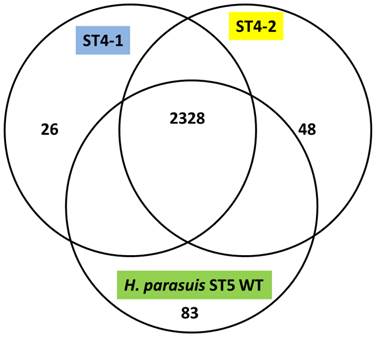
The H. parasuis serovar 4 strains (serovar 4-1 and serovar 4-2) mapped approximately 90% to the reference strain a virulent H. parasuis serovar 5 isolate, indicating a very high homology. The assemblies analyzed were approximately 190 and 150 Kb in length and consisted of 194 and 149 contigs for H. parasuis serovar 4-1 and H. parasuis serovar 4-2 respectively to cover the entire genome. We obtained 255x, 300x and 200x coverage for virulent H. parasuis serovar 4-1, H. parasuis serovar 4-2 strains and H. parasuis serovar 5 WT respectively. The comparison criteria set for all the three strains was that open reading frames (ORFs) having <80% overall base coverage were called absent. Among 2456 ORFs identified, approximately 919 were hypothetical proteins and 2328 ORFs were common to all the three strains. There were 83 genes unique to the virulent H. parasuis serovar 5, 26 genes unique to H. parasuis serovar 4-1, and 48 genes unique to H. parasuis serovar 4-2 (Figure 1). The list of individual genes unique to each isolate is summarized in Supplementary Material: Table 2. Both H. parasuis serovar 4 strains lacked enzymes such as UDP-glucose-4-epimerase, 'toxin-antitoxin system” and a large number of hypothetical proteins which were present in H. parausis serovar 5. However, the genes involved in altering the outer membrane protein (OMP) and lipooliosaccharide (LOS) structure are sialyltransferase, glycosyl transferases, polysaccharide biosynthesis protein capD, spore coat polysaccharide biosynthesis protein C and polysaccharide export protein were present and analyzed. These proteins are involved in serovar determination, immune evasion and act as virulence factors. In addition, H. parasuis serovar 4-1 and serovar 4-2 contain genes which encode other virulence and virulence associated factors found in the “wild type” H. parasuis serovar 5. The genomes of H. parasuis serovar 4 strains have been deposited at DDBJ/EMBL/GenBank® under the accession numbers JJNQ00000000 (serovar 4-1), JJNR00000000 (serovar4-2). The version described in this paper is JJNQ00000000.1 and JJNR00000000.1.
Venn diagram representing the number of genes unique to H. parasuis Sserovar 4-1, SEROVAR 4-2 and H. parasuis serovar 5 wild type.
Sialyltransferase
Sialylation of LOS has been implicated as a bacterial virulence factor involved in inhibiting antibody binding and enhancing bacterial serum resistance (16, 17). The genes encoding α-2, 3-sialyltransferases are involved in LOS biosynthesis in pathogenic members of the Pasteurellaceae family such as Neisseria gonorrheae, Haemophilus ducreyi, Haemophilus influenzae and Histophilus somni (17,18). Sialylated LOS aids host-immune evasion by molecular mimicry. Some pathogenic bacteria also utilize linkage-specific sialyltransferases like, α-2, 3 linkages to galactose, or α-2, 6 linkages to galactose or N-acetylgalactosamine to sialylate the LOS to avoid host innate defense mechanisms (17). Sialylated LOS have been reported to induce significantly less cytokine response from macrophages by inhibiting the TLR-4 signaling pathway, which in turn induces significantly less transcription factor NF-κB than de-sialylated LOS (19,20).
Sialyltransferase-encoding gene (lsgB) is known to be present only in virulent H. parasuis strains, which also display an increased resistance to serum-mediated killing and phagocytosis by alveolar macrophages than strains lacking lsgB (21). However, pigs infected with variant H. parasuis serovar 4 exhibited signs of acute pneumonia and septicemia indicating a possible “cytokine storm” as a result of infection. Furthermore, in piglet challenge experiments, H. parasuis serovar 4 was the dominant species recovered from the tissue samples. When mice were injected with a de-sialylated H. parasuis serovar 4 mutant all the mice died within 48 h due to septicemia, indicating an exuberant immune response. In contrast, mice injected with sialylated (wild type) H. parasuis serovar 5 did not die, but had injection site swelling. Providing further evidence for the lack of involvement of the sialyltransferase gene in virulence and evidence that some H. parasuis serovar 4 strains may have alternate pathways for adding sialic acid to proteins and polysaccharides.
Haemophilus parasuis serovar 4 and serovar 5 lack specific de novo sialic acid biosynthesis genes, but the genome encodes two tripartite ATP-independent periplasmic transporters, Neu5Ac TARP (HPM_1229, HPM_0026), TRAP permease (HPM_0027) and CMP-Neu5 Ac synthetase gene (HPM_0608), acylneuraminate cytidylyltransferase (HPM_1673) indicating the presence of a sialic acid scavenge mechanism similar to that of H. influenza and synthesis of CMP-Neu5Ac (17). Since the variant H. parasuis serovar 4 lacks α-2, 3-sialyltransferases, we hypothesized that the amount of sialic acid coupled to LOS should be less than serovar 5. The amount of sialic acid coupled to LOS was determined after treating the cells with neuraminidase and the H. parasuis serovar 4 strains had significantly less sialic acid residues linked to LOS when compared to serovar 5 WT (Fig 2).
Amount of sialic acid released from H. parasuis serovar 5 WT and serovar 4 after neuraminidase treatment.
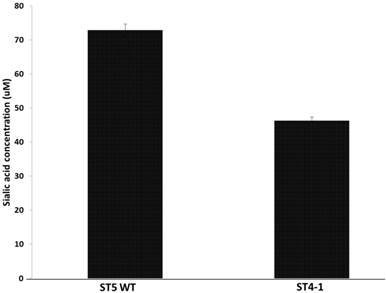
These results contradict the current thinking that sialic acid is necessary for H. parasuis virulence. The variant H. parasuis serovar 4 strains possess other virulence factors that help subvert host-immune response and aid in pathogenesis.
Glycosyl transferases
The variant H. parasuis serovar 4-1 and serovar 4-2 strains lack four glycosyl transferases (HPM_1373, HPM_1372 and HPM_1370 and HPM_1371). Glycosyl transferases catalyze the reaction involving the addition of carbohydrate side group to a protein or lipid molecule. Addition of carbohydrate residues to proteins significantly alter biophysical and structural properties such as the solubility index, resistance to proteases, stability, protein-protein interactions and most importantly the immunogenicity of pathogenic bacteria (22-26).
The lack of glycosyl transferases among the variant H. parasuis serovar 4 strains suggests the existence of an alternate pathway for glycosylation as different groups of glycosyl transferases have overlapping functions. Many enzymes involved in cell wall biosynthesis like N-acetylmuramyl-tripeptide synthetase protein (HPM_1513), MurNAc-pentapeptide synthetase (HPM_1514), phospho-N-acetylmuramoyl-pentapeptide-transferase (HPM_1515), UDP-N-acetylmuramoyl-L-alanyl-D-glutamate synthetase (HPM_1945), D-alanine-D-alanine ligase (HPM_2304), UDP-3-O-acyl-N-acetylglucosamine deacetylase (HPM_2413) and exopolysaccharide biosynthesis protein (HPM_0770) are identical across all three strains. A complete list of serovar 4 genes is provided in Supplementary Material: Table 3. Modified LOS and OMP glycosylation pattern of variant H. parasuis serovar 4 strains most likely help the bacterium avoid phagocytosis.
Polysaccharide export protein and polysaccharide biosynthesis protein CapD
Polysaccharide export protein and polysaccharide biosynthesis protein are located as contiguous ORFs in H. parasuis serovar 4 and serovar 5 strains. Bacteria export extracellular polysaccharides (EPS) and capsular polysaccharides (CPS) through complex network of transport systems (27). These biopolymers have various biological functions, such as providing reserve material or as part of the cell protective structure, and can provide an advantage for pathogenic bacteria under certain environmental conditions.
In pathogenic bacteria, production and export of EPS and CPS poses a major challenge as these high-molecular-weight hydrophilic polymers must be assembled and exported across the envelope, without compromising the essential functional properties of the envelope (27). Two major pathways with different polymer biosynthesis strategies are employed by bacteria in the assembly of most EPS/CPS: the Wzy-dependent and ATP-binding cassette (ABC) transporter-dependent pathways (28). They converge in an outer membrane export step mediated by a member of the outer membrane auxiliary (OMA) protein family. Outer membrane auxiliary proteins form outer membrane efflux channels for the biopolymers. The polysaccharide co-polymerase (PCP) family of enzymes interacts with OMA proteins forming a trans-envelope scaffold for polymer export (28). The variant H. parasuis serovar 4 strains have numerous ABC transporter systems similar to virulent H. parasuis serovar 5 but lack the polysaccharide export protein (HPM_0299) wza which reveals a functional overlap between ABC transporters in exporting complex carbohydrates and biopolymers.
The capD gene (HPM_0300) encodes a polysaccharide biosynthesis protein which has been implicated in H. parasuis virulence. However, a characteristic feature of this gene that is associated with the pathogenicity of H. parasuis is not understood (29). This domain is found in diverse bacterial polysaccharide biosynthetic proteins including the WalL protein, mannosyl-transferase and epimerases (30). The CapD protein is required for biosynthesis of type 1 capsular polysaccharide in Staphylococcus spp. (31) and serum-resistance in H. parasuis (32). Despite lacking this domain the variant H. parasuis serovar 4-1 and serovar 4-2 strains are highly virulent and were recovered in pure cultures from experimentally infected dead pigs, possibly supporting serum-resistance and virulence without the capD protein.
Earlier reports indicate that the deletion of capD gene significantly attenuates H. parasuis serotype 5 SH0165 (GenBank No. CP001321.1) pathogenicity and the complementation of this gene restored the pathogenicity in piglets (32). In addition, the capD deleted H. parasuis serotype 5 SH0165 strains were not recovered from challenged piglets. However, both H. parasuis serotype 5 SH0165 wild type and complemented-capD strains were recovered from most of the systemic sites (32). The variant H. parasuis serovar 4 strains caused a rapid onset of clinical signs and death (50% mortality) in experimentally infected pigs. Necropsy of the dead pigs revealed lung lesions typical of H. parasuis serovar 4 and isolation of the original challenge strain.
PCR analysis of virulence genes
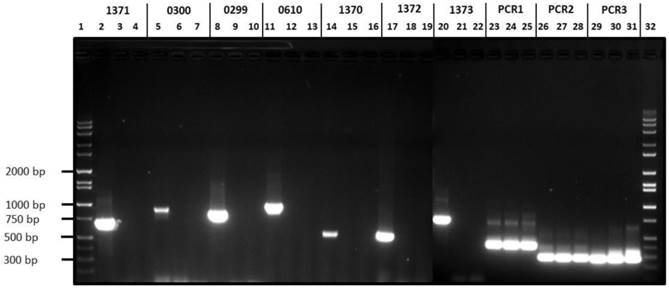
The variant H. parasuis serovar 4-1 and serovar 4-2 strains lack ORFs HPM_1370, HPM_1371, HPM_1372, HPM_1373, HPM_0299 and HPM_0300 reiterating our in silico based analysis (Fig 3). On the other hand, H. parasuis variant serovar 4 strains possess all the three virulence associated trimeric autotransporters (vtaA) domains similar to virulent H. parasuis serovar 5. Comparative genomic analyses revealed that group 3 vtaA is highly conserved among the invasive and non-invasive strains, while groups 1 and 2 vtaA were detected only in the virulent strains (33). However all three domains were detected in highly virulent serovar 4-1, serovar 4-2, and serovar 5 (Fig. 3) as well as in an avirulent serovar 5 strain (data not presented). We speculate that characterization of avirulent and virulent H. parasuis strains based of vtaA domains may not be a reliable approach given the complex gene regulation mechanism involved in determining H. parasuis virulence.
Agarose gel electrophoresis of H. parasuis serovar 4-1, serovar 4-2 and H. parasuis serovar 5 wild type using primers listed in Table 1. Lanes 1 & 32, Molecular weight markers; Lanes 2, 5, 8, 11, 14, 17, 20, 23, 26 & 29, PCR products from serovar 5 WT; Lanes 3, 6, 9, 12, 15, 18, 21, 24, 27 and 30, PCR products from serovar 4-1; Lanes 4, 7, 10, 13, 16, 19, 22, 25, 28 & 31, PCR products from serovar 4-2.
Other virulence and virulence associated genes
In addition to the known virulence genes, H. parasuis serovar 4 and serovar 5 genomes encode a large number of other virulence and virulence associated genes. These include regulation of uptake of metal ions, MerR family transcriptional regulators, macrophage infectivity potentiator-related protein, hemolysin, opacity associated protein, toxin antitoxin system, colicins, cytolethal distending toxin and virulence associated protein D. The complete lists of genes identified in variant H. parasuis serovar 4 strains are listed in Supplementary Material: Table 3.
Regulation and uptake of metal ions
Like many pathogenic bacteria, the variant H. parasuis serovar 4-1, serovar 4-2 strains and serovar 5 have extensive regulatory and protein-coding systems exclusively devoted to maintaining the homeostasis of biologically required metal ions, most of which are acquired from the environment. Of all the metal acquired ions, iron is the primary regulator of many virulence genes. Our findings reveal that the variant H. parasuis serovar 4-1, serovar 4-2 and serovar 5 have an extensive protein network to regulate iron uptake. Iron is required by a number of essential metabolic enzymes, including the cytochromes, ribonucleotide reductase, Fe-S cluster biogenesis and activation/regulation of virulence genes but is often an in vitro growth limiting factor for pathogenic bacteria due to its low solubility in water at neutral pH (34).
Mammalian hosts lack free iron for uptake, since most of it is stored in intracellular or extracellular tightly bound forms like transferrin, lactoferrin, haemopexin and haptoglobin (34, 35). Many bacterial outer membrane (OM) receptor proteins like ferroxamine (HPM_2297) bind to lactoferrin or ferritin prior to mediating internal transport. These receptors are usually 22-stranded β-barrel proteins containing extracellular loops which bind substrates, and an N-terminal region or plug, that folds into the barrel near the periplasmic surface (36). Since there are no ionic gradients to drive this transport, transport across the OM is coupled to the proton motive force of the cytoplasmic membrane via a periplasm-spanning complex like TonB, ExbB and ExbD (37). The H. parasuis periplasm-spanning complex is composed of TonB (HPM_0089 and HPM_0090), ExbB (HPM_0091), and ExbD (HPM_0092). Once in the periplasm, the uptake of Fe (III) or Fe (III)-chelate occurs through the transmembrane channel of ATP-binding cassette (ABC) transporters found in the plasma membrane, in a process mediated by ATP hydrolysis (38). Haemophilus parasuis serovar 4 and serovar 5 also possess three heme binding proteins (HPM_0689 HPM_2386, HPM_0785) which bind to heme and hemin and are involved in iron acquisition. Variant H. parasuis serovar 4 also encodes iron-binding protein IscA (HPM_1829), which transfers intracellular free iron to iron-sulfur clusters to apo-ferredoxin, iron utilization protein hugX (HPM_0784), ferric iron reductase (HPM_1543), involved in ferric hydroximate transport, very high affinity iron scavenging proteins hemopexin B and A (HPM_ 1235, HPM_1236), two periplasmic iron uptake and binding protein (HPM_1161 and HPM_1150), ferric transporter ATP-binding subunit (HPM_1466), high affinity heme/hemopexin utilization protein C/outer membrane receptor protein to transport Fe ions (HPM_1031) and ferric uptake regulation protein (HPM_1051). The heme exporter protein B (HPM_2191) which exports heme to the periplasm for the biogenesis of c-type cytochromes is adjacent to cytochrome c biogenesis ATP-binding export protein CcmA (HPM_2192).
Haemophilus parasuis serovars 4 and serovar 5 encode two outer membrane TonB dependent receptors or iron regulated outer membrane proteins, IROMP (HPM_0168, HPM_0415). Encoded in H. parasuis serovar 5 and serovars 4 are three small multifunctional proteins, frataxin (HPM_1028), ferredoxins (HPM_0902) and Ferritin/ DNA-binding stress protein (HPM_2074). Frataxin acts as an iron chaperone during cellular heme and iron-sulfur (Fe-S) cluster production, scavenges iron and stores iron during iron overload, repairs oxidatively damaged aconitase Fe-S clusters, reduces oxidative stress by moderating the concentration of reactive oxygen species (ROS) and is involved in energy conversion and oxidative phosphorylation (39-42).
Thus, H. parasuis serovar 4-1, serovar 4-2 and serovar 5 genomes encode a large number of proteins with overlapping functions involved in maintaining iron homeostasis, indicating the critical role of iron in H. parasuis pathogenesis. These proteins could also possibly be targeted to develop universal vaccines or for strain identification.
In addition to iron, H. parasuis requires metal ions like molybdenum, zinc, copper, nickel, and magnesium for the activation of many metallo-proteases, metabolic enzymes and DNAse. H. parasuis serovar 4-1, serovar 4-2 and serovar 5 has an extensive network of proteins to acquire these metals which include molybdenum cofactor biosynthesis (HPM_ 0200), molybdoprotein biosynthesis protein, molybdoprotein biosynthesis MoeB and two MoeA (HPM_0201, HPM_1239 and HPM_1012), moylbdate ATP transporter (HPM_1718), molybdate ABC transporter (HPM_1719), molybdenum transport protein (HPM_0692), molybdate-binding periplasmic protein (HPM_0693), molybdenum cofactor biosynthesis protein C' (HPM_0289), molybdopterin converting factor (HPM_0290). In addition, zinc ABC transporter (HPM_0361, HPM_1198), high-affinity zinc transporter periplasmic component (HPM_0485), zinc/copper dismutase (HPM_0873), copper homeostastsis (HPM_1035), nickel transport permease (HPM_2246), nickel-binding periplasmic precursor protein, nikA (HPM_0691), ABC-type nickel/cobalt efflux system, permease component (HPM_2454), Mg2+/Co2+ transporter (HPM_2352), magnesium dismutase (HPM_1717), and magnesium transport corA protein (HPM_ 0841).
MerR family of transcriptional regulators
The regulation of metal ion homeostasis is exclusively controlled by MerR family of transcriptional regulators (43). MerR family of regulators have been exclusively found in eubacteria but not yet identified among archaebacteria or eukaryotes (43). MerR transcriptional regulators function nearly exclusively as transcriptional activators of gene expression, and are required for metal efflux or detoxification, defense against oxidative, biotic or abiotic stress and provide drug resistance (44).
In H. parasuis serovars 4 and serovars 5, MerR regulator (HPM_1347) is a 15.7 kDa protein and belong to a helix-turn-helix (HTH) MerR-SF superfamily. Haemophilus parasuis MerR is similar to other bacterial MerR regulatory proteins and follow a canonical N-terminal HTH-DNA binding domain which is about 40 amino acids long and a C-terminal metal-coordinating domain (between 80-130 amino acids) that is specific to the metal ion recognized (45, 46). The MerR families of metal-binding and metal-responsive proteins are unique in that they activate transcription from unusual promoters with a high spacing of 19 bp between the-35 and-10 sequences (45). These cytoplasmic transcription factors bind metal ions through cysteine or histidine residues (46). Although different bacteria and have a common design i.e. conserved primary structure, they can effectively discriminate metals in vivo (47). Based on NCBI database (http://www.ncbi.nlm.nih.gov/Structure/cdd/wrpsb.cgi?RID=1A9YCPJB014&mode=all) analysis (E-value: 7.89e-39) H. parasuis- Mer R regulator appears to be a heavy metal (copper, cadmium, lead, zinc) resistance transcription regulator.
The presence of these extensive metal acquisition genes along with a specific transcriptional regulator exhibits a very complex gene regulation mechanism involved in the expression of H. parasuis virulence genes.
Macrophage infectivity potentiator-related protein
Macrophage infectivity potentiator (MIP)-related proteins have been identified in many pathogenic bacteria such as Neisseria, Actinobacillus, Legionella, Chlamydia and in faculatively intracellular pathogens like Coxiella burnetii and Burkholderia pseudomallei and the protozoan parasite, Trypanosoma cruzi (48-51). Variant H. parasuis serovars 4 and serovar 5 genomes encode a putative macrophage infectivity potentiator-related protein (HPM_0216), which is a 20 kDa outer membrane protein. Our analyses (http://blast.ncbi.nlm.nih.gov/Blast.cgi) shows that H. parasuis serovars 4 and serovar 5 MIP belong to the carboxymuconolactone decarboxylase (CMD) superfamily (COG2128), exhibiting a multidomain architecture similar to other pathogenic bacteria.
Haemophilus parasuis serovar 5 and serovars 4 MIP contain overlapping alkylhydroperoxidase AhpD family core domain, alkylhydroperoxidase domain protein, Avi_7169 family and a gamma-carboxymuconolactone decarboxylase. Like the rest of the prokaryotic MIP-like proteins, the predicted N- terminal structure of H. parasuis serovars 4 and serovar 5 MIP contain three large alpha-helices followed by a short beta-sheet or turn.
Burkholderia pseudomallei MIP-like protein is sensitive to immunosuppressants like FK506 and rapamycin which abolish its peptidylprolyl isomerase activity. Burkholderia pseudomallei mutants lacking MIP exhibit reduced ability to survive within cells and are significantly attenuated in vivo (50). In addition, MIP plays a vital role in Chlamydia spp. entry into McCoy cells (48). A similar function for H. parasuis MIP cannot be ruled out and may aid in intracellular invasion and survival of the bacteria. MIP-like proteins could potentially serve as a target for universal vaccine.
Hemolysin
Hemolysins are toxic to erythrocytes, but some bacterial species produce hemolysins which lyse leukocytes as well (52-56,59). The variant H. parasuis serovar 4-1, serovar 4-2 and serovar 5 have two hemolysin activation/secretion proteins (HPM_2302, HPM_1788) and two hemolysin structural proteins (HPM_1789, HPM_2290). The ORFs HPM_1788 and HPM_1789 are adjacent to each other, similar to those found in H. ducreyi (hhdB and hhdA), Serratia marcescens (shlB and shlA), Proteus mirabilis (hpmA and hpmB) and Edwardsiella tarda. However, ORFs HPM_ 2302 and HPM_2290 are not adjacent. In addition H. parasuis serovars 4 and serovar 5 genomes also encode AphA-like protein/membrane protein affecting hemolysin expression (HPM_2138), a 21 kDa hemolysin precursor protein (HPM_0599) and another possible hemolysin structural protein (HPM_0082). Earlier studies have shown that the S. marcescens, ShlB is an outer membrane protein which is required for secretion and activation of the hemolysin structural protein, ShlA (57,58). Once secreted, ShlA interacts with target cell membranes, oligomerizes, and forms pores on the cell membrane resulting in target cell lysis similar to other members of the “RTX” toxin family indicating a similar function in H. parasuis.
Opacity associated protein
Opacity-associated proteins (OapA and OapB) have been implicated in phase variation, adhesion to epithelial cells during initial colonization and cell invasion in H. influenza, and Neisseria gonorrheae (60, 61). H. parasuis serovar 5 and serovars 4 strains encode two OapA and OapB proteins (HPM_ 0924, HPM_0925) which help in the initial colonization and adhesion along with fimbriae/adhesins and pili complex loci ( HPM_0366, HPM_0367, HPM_0368, HPM_0371, HPM_1452, HPM_1455 and HPM_1637).
Toxin-antitoxin system
Haemophilus parasuis serovars 4 and serovar 5 genomes encode numerous toxin-antitoxin systems (TAS) which are abundant, diverse, horizontally mobile genetic elements (62-64). Toxin-antitoxin systems are frequently limited to bacterial and archaeal genomes and are involved in a variety of functions including plasmid stabilization, transcription regulation, enhancing resistance mechanisms and RNA-interference (63-65). Haemophilus parsuis serovars 4 and serovar 5 TAS help prevent hostcell-death by a group of protein/antitoxins of the toxin-antitoxin stability system (HPM_1007). These include the antitoxin/toxin system zeta toxin, signal recognition particle GTPase protein (HPM_1145), fic family toxin-antitoxin (HPM_1182), addiction module antitoxin/putative RelE toxin-like protein, plasmid stabilization system (HPM_1184), toxin component, RelE family (HPM_0312), transcriptional regulator/antitoxin, MazE protein (HPM_1226), antitoxin ChpS/ transcriptional regulator/antitoxin, MazE/ putative plasmid stable inheritance protein (HPM_1862), growth inhibitor, PemK-like, autoregulated/transcriptional modulator of MazE/toxin, MazF, plasmid stable inheritance protein K protein (HPM_1863), plasmid stability protein StbD, (HPM_2272), HicA and B (HPM_2185, HPM_0011 and HPM_0012). The number of modules in a TAS may range from one to eight (66,67). However, most of the regulated TAS are two-component systems and function similar to HicA and HicB.
The ORFs HicA and B (HPM_0011 and HPM_0012) are contiguous and attached to competence protein comM (HPM_0013), whereas another HicA (HPM_2185) is associated with an endonuclease (HPM_2184). The HicB protein has a partially degraded RNAse H fold, whereas HicA has a double-stranded RNA-binding domain (68). The stable combination of these two domains suggests transcription regulation mediated through RNA, most likely by RNA binding-degradation (RNA interference, RNAi) mechanism. In most HicB proteins, the RNAse H-like domain is fused to a DNA-binding domain, of the ribbon-helix-helix or to the helix-turn-helix motif (68). TheTAS proteins containing these DNA-binding domains function as antitoxins (68). Haemophilus parasuis serovars 4 and serovar 5 strains have numerous TAS genes located within their pathogenicity islands that are amenable for horizontal transfer and function as accessories to chief virulence genes similar to pathogenic bacteria.
Virulence associated protein D
Haemophilus parasuis serovars 4 and serovar 5 genomes encode virulence associated protein D (VapD), (HPM_1572) which shares 100% identity to VapD of a serovar 5 H. parasuis SH0165 (YP_002476324), H. parasuis ZJ0906 (YP_008124455) and H. parasuis SW114 (EQA00157.1). However, they are only 90% identical to Actinobacillus minor (WP_005826028), 64% to Neisseria lactamica (WP_003711106.1), 63% to N. meningitides (WP_002238271.1), and 62 % to CRISPR associated Cas2 family protein of N. meningitides (WP_002251489).
Rhodococcus equi contains a highly characterized vapD gene found on an 80-90 kb virulence plasmid which contains a 27.5 kb pathogenicity island encoding a family of seven Vap proteins including VapA-G (69, 70). Rhodococcus equi mutants lacking 7.9 kb DNA region spanning five vap genes (vapA, -C, -D, -E and -F) are avirulent and are rapidly cleared by the mouse immune system compared to wild type (70). Furthermore, R. equi isogenic plasmid-cured mutant strains lose their ability to survive in alveolar macrophages and fail to induce pneumonia in foals (71,72). The vap genes are also induced by H2O2 (73). The VapD protein along with macrophage infectivity potentiator may also be involved in initiating and maintaining H. parasuis intracellular infection, especially within macrophages. Therefore, these antigens could be incorporated in a novel vaccine development strategy.
Colicin
Colicins are heat-labile proteins first identified in certain strains of E. coli that harbor a single colicinogenic plasmid (74). Since this report, colicins have been identified in many bacterial strains which are antagonistic to other closely related strains. Colicinogenic strains of pathogenic bacteria are widely distributed in nature and are particularly abundant in the animal intestinal tracts (75).
Haemophilus parasuis serovars 4 and serovar 5 colicins, like other bacteria, are not synthesized under normal conditions since the colicin operon is repressed by the LexA protein (HPM_1027). Haemophilus parasuis serovars 4 and serovar 5 encode colicin V (HPM_1084) and a colicin transport protein TolQ (HPM_1307) conferring a competitive advantage to strains occupying nasal cavities and upper respiratory tracts of the pigs.
Cytolethal distending toxins
Cytolethal distending toxins (CDT) are heterotrimeric toxins and have been identified in various Gram-negative bacteria including Campylobacter spp, E. coli, H. ducreyi, A. actinomycetemcomitans, S. dysenteriae, and Helicobacter spp. (76-81). Upon entry into the target cells, CDTs randomly nick cellular DNA leading to apoptosis (82). These toxins also trigger G2/M cell cycle arrest in specific mammalian cell lines leading to enlargement of distended cells that result in necrosis (83). This toxin is internalized via the Golgi complex and transported to the endoplasmic reticulum in a retrograde fashion similar to Cholera Toxin A1 (84). Cytolethal distending toxin production is dependent on the expression of three (cdtA, cdtB, and cdtC) contiguous genes in the operon (85, 86).
Haemophilus parasuis H. serotypes 4 and H. serotype 5 genomes contain one gene encoding CDT internalizing protein, cdtA (HPA_2217), but lack the active toxin cdtB and an internalizing protein, cdtC. The possible reasons for missing cdtB and cdtC genes are, either due to a gap in the map-based assembly or they may truly lack a functional CDT operon. If this is the case, H. parasuis would have similarities to Campylobacter spp. strains lacking CDT that cause enteric diseases in human (87).
Unmapped genes
Analysis of the unmapped regions yielded a total 263 genes for serovar 4-1 and 323 genes for serovar 4-2 (Supplementary Material: Tables 4 and 5). Open reading frames (ORFs) less than 300 bp are usually repeats and often do not encode functional proteins. Therefore a cut off value of 300 bp was set and ORFs ≥300 bp were mapped and annotated. This resulted in 190 ORFs for serovar 4-1 and 237 ORFs for serovar 4-2 respectively. The unmapped regions of serovar 4-1 and serovar 4-2 contain horizontal gene transfer elements, hypothetical proteins, transcription factors, phage genomes, transporters, housekeeping genes (cell wall/LPS biosynthesis) and a toxin antitoxin protein zeta toxin. However, the uniqueness of these genes to each strain could not be verified since this was a map-based assembly on draft genomes.
In summary, the comparative genomic analysis presented here clearly indicates the variant H. parasuis serovar 4-1 and serovar 4-2 strains have extensive LOS and OMP structural modifications that aid in nasal colonization, host immune evasion and possibly serum-resistance. In addition, the lack of sialic acid residues on serovar 4-1 and serovar 4-2 LOS may lead to a “cytokine storm” resulting in septicemia, pulmonary edema and severe pneumonia as evidenced by the lung pathology. When piglets were challenged with variant H. parasuis serovars 4, 50% died within 48 h indicating the highly virulent nature of the variant serovar 4 strains. The variant H. parasuis serovar 4 strains also harbors a more diverse repertoire of virulence associated genes than previously described. The combination of capsule modification and phase variation due to LOS substitutions could help variant H. parasuis serovar 4 escape host immune system and possibly transform into a deadly pathogen and lead to a disease outbreak.
Methods
Bacterial isolates and genomic DNA sequencing
The H. parasuis serovar 4 (strains were isolated by the Newport Laboratories and dubbed as H. parasuis serovar 4-1 and H. parasuis serovar 4-2. Haemophilus parasuis was cultured on Trypticase Soy Agar (TSA) plates supplemented with 5% bovine serum and 10 mg/ml nicotinamide adenine dinucleotide (NAD) at 37ºC in 5% C02 for 36 hours. Following incubation the bacterial cells were removed from the plate with sterile PBS and washed twice with PBS prior to DNA extraction.
Genomic DNA from H. parasuis serovar 5 wild type (Bioproject accession PRJNA218837) and serovar 4 strains was extracted using bacterial genomic DNA isolation kit (Edge Biosystems, Gaithersburg, MD, USA). The quality of the genomic DNA was determined by PicoGreen (Life Technologies, Grand Island, NY, USA). The genomic DNA library was generated using a TruSeq® Library preparation kit (Illumina Inc., San Diego, CA, USA) following the manufacturer's recommendation and sequenced on Illumina HiSeq 2000 to obtain on 50 bp PE reads. Whole genome sequencing was performed at the University of Minnesota Genomics Center. Allll the H. parasuis strains isolated in this study were confirmed by the Department of Veterinary Medicine, University of Montreal, Montreal, Canada.
Sequence Analysis
Haemophilus parasuis serovar 5 GenBank isolate SH0165 (GenBank accession number CP001321.1, 88) was used as the reference genome to assemble and map H. parasuis serovar 5(Bioproject accession PRJNA218837). Concatenated genome of H. parasuis serovar 5 was used to perform comparative genomic analysis H. parasuis serovar 4-1 and serovar 4-2 variants. The web based software ABACAS (http://abacas.sourceforge.net/) was used to align the SH0165 contigs generated and identify putative gap sizes, orientation, and order of contigs. A custom Python script was used to concatenate contigs, fill the gaps with N's, and transfer the annotations. Additional annotations were added to the resulting sequence to highlight gaps, and to label the original contigs. The contigs which could not be aligned against the reference genome were included at the end of the concatenated sequence.
The reads were filtered for quality and Illumina TruSeq adapters using an in-house script which removed reads with 5 or more bases that had a PHRED Quality value <30. Because of very high coverage, duplicate reads were expected and reads were only analyzed if at least two duplicates existed. A maximum of 10 duplicates were kept for each read. Following quality filtering, reads were mapped against SH0165 and a concatenated set of contigs was aligned and oriented using SH0165 as a reference.
Consensus GenBank formatted sequences for each serovar 4 isolate were generated by mapping reads against serovar 5 WT (Bioproject Accession: PRJNA218837) using Bowtie 2 and then calling a mapping consensus using Samtools. Annotations from serovar 5 WT were then copied to the new serovar 4 consensus sequence. Reads were mapped for each isolate using Bowtie2 v 2.0.6 (http://bowtie-bio.sourceforge.net/bowtie2/index.shtml) and default primers against H. parasuis serovar 5. . Filtering was then carried out to remove multiple mapped reads and MAPQ <10 using a custom script. Based on the mapping results, SAMtools (http://samtools.sourceforge.net/ ) was employed to generate the consensus sequences. This sequence data was annotated using the reference sequence and formatted as a GenBank file for Artemis compatibility using custom scripts and BioPython. (http://biopython.org/wiki/Biopython).
The reads which could not be mapped to the GenBank reference strain, by Bowtie2, were identified and assembled using the Newbler v2.6® software (GS de novo Assembler, Roche). The gene prediction program Prodigal v2.6 (http://prodigal.ornl.gov/) was used to predict genes from the assembled contigs. Putative genes were assigned anannotation using tool BLANNOTATOR (http://www.biomedcentral.com/1471-2105/13/33).
Mapping coverage was calculated for each position in reference using SAMtools. Gene annotation information was used to calculate the percentage of bases which had coverage for each gene. Genes in which less than 80% of bases had coverage were called absent and classified as unmapped genes.
Finally, unmapped reads were assembled using Newbler v2.6®, putative genes determined using Prodigal, and annotated using the software tool BLANNOTATOR (http://www.biomedcentral.com/1471-2105/13/33).
PCR Analysis of Capsule and Putative Virulence Genes
Genomic DNA from all the 3 strains was extracted using the bacterial genomic DNA isolation kit (Edge Biosystems, Gaithersburg, MD, USA). Polymerase chain reaction (PCR) assay's specific for H. parasuis serovar 4 and serovar 5 genes were performed using the set of primers described in Table 1. Polymerase chain reaction assays were carried out in a final, 25 µl volume with GoTaqH PCR SuperMix® (Promega Inc., Madison, WI, USA) with 0.2 mM of each primer and 10 ng of genomic DNA from each isolate respectively. The PCR cycling conditions consisted of an initial denaturation at 95 ºC for 5 min followed by 35 cycles of denaturation at 95 ºC for 30 sec, annealing at 55 ºC for 30 sec, and extension at 68 ºC for 1 min, and a final elongation at 72 ºC for 5 min. All PCR products were visualized after electrophoresis in 1.0% agarose gels run at 7.0 V/cm by staining with ethidium bromide.
List of primer pairs used to amplify virulence and virulence associated genes from H. parasuis serovar 4-1, serovar 4-2 and H. parasuis serovar 5 wild type.
| Primers | Sequence (5'→3') | Product (bp) | Amplicon |
|---|---|---|---|
| 0610 FOR | ATGAATTTGATTATTTGTATGACTCCATTTC | Sialyltransferase | 969 |
| 0610 REV | CTATTGGCATGTGTAGTCAATTACTTC | ||
| 0300 FOR | ATGTTAATGCCATTAATTTATTCATTG | CapD protein | 780 |
| 0300 REV | TCGAACCGATAGAACCAGCAGCACCAGTC | ||
| 0299 FOR | ATGTGTAAGTTAACTAAAGCTCTTG | Polysaccharide export protein | 840 |
| 0299 REV | AGCAATTGCTTCGGTTAACGTCATAC | ||
| 1370 FOR | ATGCTAAAAAGAGTGTTTGATATTTTC | Glycosyl transferase | 540 |
| 1370 REV | TATATTATGATTAACATAATC | ||
| 1371 FOR | ATGAACTTTCTACCATTCGCCCTTCCCG | Glycosyl transferase | 520 |
| 1371 REV | ATTATATTTGAATCCAGGTTCAATG | ||
| 1372 FOR | ATGAAATTGTCTGTCTTAATGGCTGT | Glycosyl transferase | 720 |
| 1372 REV | TCCGCCAAATGTACATCATCAC | ||
| 1373 FOR | ATGAAATTGTCTGTCTTAATGGCTGT | Glycosyl transferase | 462 |
| 1373 REV | CTCTCATACCATACCCCAACTCAGG | ||
| YADAF1* | TTTAGGTAAAGATAAGCAAGGAAATCC | vta (PCR1) | 406 |
| PADHR1* | CCACACAAAACCTACCCCTCCTCC | ||
| YADF2* | AGCTTATATTCTCAGCACAAGGTGC | vta (PCR2) | 294 |
| PADHR2* | CCACTGATAACCTACCCCCACAGAG | ||
| YADAF3* | AATGGTAGCCAGTTGTATAATGTTGC | vta (PCR2) | 293 |
| PADH3* | CCACTGTAATGCAATACCTGCACC |
* Olvera A, Pina S, Macedo N, Oliveira S, Aragon V, Bensaid A. Identification of potentially virulent strains of Haemophilus parasuis using a multiplex PCR for virulence-associated autotransporters (vtaA). Vet J. 2012 191(2):213-218.
Sialic acid assay
Haemophilus parasuis cultures were grown as described above and heat inactivated. Briefly the bacteria removed from the culture plate, harvested by centrifugation (8,000 x g for 10 min) and washed three times with phosphate-buffered saline (1X PBS), pH 7.2, to remove secreted proteins and resuspended in PBS. Serial tenfold dilutions were made and plated on Trypticase Soy Agar plates to determine the number of colony forming units (CFU) per ml. Bacterial suspensions were heat inactivated at 60°C for 1 h and plated again to confirm heat inactivation. The cell suspension was adjusted to1010 CFU/ml. Fifty microliter aliquots of the inactivated bacterial cultures were treated with 0.5 U of neuramidinase (Sigma, St. Louis, MO, USA) and incubated for an hour at 37˚C before proceeding with the sialic acid quantitation assay (15). The sialic acid assay was performed using QuantiChrom® sialic acid assay kit (BioAssay Systems, Hayward, CA, USA) following the manufacturer's recommendations. Sialic acid standards provided with the kit were used to derive a standard curve and the amount of sialic acid released from lipooligosaccharide (LOS) was expressed as µM.
Supplementary Material
Tables 2-5.
Acknowledgements
We thank Sam Hunter, University of Idaho for analyzing the three H. parasuis genomes. We also thank Brittany Wiener, Eric Bumgardner, Danielle McKeown, Kristina Breuker and John Porth, Newport Laboratories for technical support and conducting field work.
Disclaimer
This document is provided for scientific purposes only. Any reference to a brand or trademark herein is for informational purposes only and is not intended for a commercial purpose or to dilute the rights of the respective owner(s) of the brand(s) or trademark(s).
Competing interests
The authors are employed by Newport Laboratories Inc., a company involved in producing and selling swine vaccines.
References
1. Biberstein EL, White DC. A proposal for the establishment of two new Haemophilus species. J Med Microbiol. 1969;2:75-77
2. Nicolet J. Haemophilus parasuis. In Diseases of Swine. 7th edition. Edited by Leman AD, Straw BE, Mengeling WL, D'Allaire S, Taylor DJ. Iowa: Iowa State Press. 1992:526-528
3. Amano H, Shibata M, Kajio N, Morozumi T. Pathologic observations of pigs intranasally inoculated with serovar 1, 4 and 5 of Haemophilus parasuis using immunoperoxidase method. J Vet Med Sci. 1994;56:639-644
4. Peet RL, Fry J, Lloyd J, Henderson J, Curran J, Moir D. Haemophilus parasuis septicemia in pigs. Aus Vet J. 1983;60:187
5. Little TWA. Haemophilus parasuis infection in pigs. Vet Rec. 1970;87:399-402
6. Kielstein P, Rapp-Gabrielson VJ. Designation of 15 serovars of Haemophilus parasuis based immunodiffusion using heat stable antigen extracts. J Clin Microbiol. 1992;30:862-865
7. Boerlin P, Poljak Z, Gallant J, Chalmers G, Nicholson V, Soltes GA, Macinnes JI. Genetic diversity of Haemophilus parasuis from sick and healthy pigs. Vet Microbiol. 2013;167:459-467
8. Rapp-Gabrielson VJ, Gabrielson DA. Prevalence of Haemophilus parasuis serovars among isolates from swine. Am J Vet Res. 1992;53:659-664
9. Oliveira S, Pijoan C. Haemophilus parasuis: new trends on diagnosis, epidemiology and control. Vet Microbiol. 2004;99:1-12
10. Tadjine M, Mittal KR, Bourdon S, Gottschalk M. Development of a new serological test for serotyping Haemophilus parasuis isolates and determination of their prevalence in North America. J Clin Microbiol. 2004;42:839-840
11. Biberstein EL. Our understanding of the Pasteurellaceae. Can J Vet Res. 1990;54:78-82
12. Munch S, Grund S, Kruger M. Fimbriae and membranes of Haemophilus parasuis. J Vet Med B-Infect Dis Vet Pub Health. 1992;39:59-64
13. Zucker BA, Baghian A, Traux R, O'Reilly KL, Storz J. Detection of strain-specific antigenic epitopes on the lipooligosaccharide of Haemophilus parasuis by use of monoclonal and polyclonal antibodies. Am J Vet Res. 1996;57:63-67
14. Morozumi T, Nicolet J. Morphological variations of Haemophilus parasuis. J Clin Microbiol. 1986;23:138-142
15. Warren Leonard. The thiobarbituric acid assay of sialic acids. J Biol Chem. 1959;234:1971-1975
16. Steenbergen SM, Lichtensteiger CA, Caughlan R, Garfinkle J, Fuller TE, Vimr ER. Sialic Acid metabolism and systemic pasteurellosis. Infect Immun. 2005;73:1284-1294
17. Inzana TJ, Glindemann G, Cox AD, Wakarchuk W, Howard MD. Incorporation of N-acetylneuraminic acid into Haemophilus somnus lipooligosaccharide (LOS): enhancement of resistance to serum and reduction of LOS antibody binding. Infect Immun. 2002;70:4870-4879
18. Vimr E, Lichtensteiger C. To sialylate or not to sialylate: that is the question. TIM. 2002;10:254-257
19. Vimr E, Lichtensteiger C, Steenbergen S. Sialic acid metabolism's dual function in Haemophilus influenzae. Mol Microbiol. 2000;35:1113-1123
20. Severi E, Hood DW, Thomas GH. Sialic acid utilization by bacterial pathogens. Microbiology. 2007;153:2817-2822
21. Martínez-Moliner V, Soler-Llorens P, Moleres J, Garmendia J, Aragon V. Distribution of genes involved in sialic acid utilization in strains of Haemophilus parasuis. Microbiology. 2012;158:2117-2124
22. Lairson LL, Henrissat B, Davies GJ, Withers SG. Glycosyltransferases: structures, functions, and mechanisms. Annu Rev Biochem. 2008;77:521-555
23. Yan A, Lennarz WJ. Unraveling the mechanism of protein N-glycosylation. J Biol Chem. 2005;280:3121-3124
24. Marth JD. Complexity in mammalian O-linked oligosaccharide biosynthesis engendered by multiple polypeptide N-acetylgalactosaminyl-transferases. Glycobiology. 1996;6:701-705
25. Szymanski CM, Wren BW. Protein glycosylation in bacterial mucosal pathogens. Nat Rev Microbiol. 2005;3:225-237
26. Schmidt MA, Riley LW, Benz I. Sweet new world: glycoproteins in bacterial pathogens. TIM. 2003;11:554-561
27. Cuthbertson L, Mainprize IL, Naismith JH, Whitfield C. Pivotal roles of the outer membrane polysaccharide export and polysaccharide co-polymerase protein families in export of extracellular polysaccharides in gram-negative bacteria. Microbiol Mol Biol Rev. 2009;73:155-177
28. Larue K, Ford RC, Willis LM, Whitfield C. Functional and structural characterization of polysaccharide co-polymerase proteins required for polymer export in ATP-binding cassette transporter-dependent capsule biosynthesis pathways. J Biol Chem. 2011;286:16658-16668
29. Zhou H, Yang B, Xu F, Chen X, Wang J, Blackall PJ, Zhang P, Xia Y, Zhang J, Ma R. Identification of putative virulence-associated genes of Haemophilus parasuis through suppression subtractive hybridization. Vet Microbiol. 2010;144:377-383
30. Whitfield C. Biosynthesis and assembly of capsular polysaccharides in Escherichia coli. Annu Rev Biochem. 2006;75:39-68
31. Lin WS, Cunneen T, Lee CY. Sequence analysis and molecular characterization of genes required for the biosynthesis of type 1 capsular polysaccharide in Staphylococcus aureus. J Bacteriol. 1994;176:7005-7016
32. Wang X, Xu X, Wu Y, Li L, Cao R, Cai X, Chen H. Polysaccharide biosynthesis protein CapD is a novel pathogenicity-associated determinant of Haemophilus parasuis involved in serum-resistance ability. Vet Microbiol. 2013;164:184-189
33. Pina S, Olvera A, Barcelo A, Bensaid A. Trimeric autotransporters of Haemophilus parasuis: generation of an extensive passenger domain repertoire specific for pathogenic strains. J Bacteriol. 2009;191:576-587
34. Laham N, Ehrlich R. Manipulation of iron to determine survival: competition between host and pathogen. Immunol Res. 2004;30:15-28
35. Litwin CM, Calderwood SB. Role of iron in regulation of virulence genes. Clin Microbiol Rev. 1993;6:137-149
36. Ferguson AD, Chakraborty R, Smith BS, Esser L, Van der Helm D, Deisenhofer J. Structural basis of gating by the outer membrane transporter FecA. Science. 2002;295:1715-1719
37. Braun V, Gaisser S, Herrmann C, Kampfenkel K, Killmann H, Traub I. Energy coupled transport across the outer membrane of Escherichia coli: ExbB binds ExbD and TonB in vitro, and leucine 132 in the periplasmic region and aspartate 25 in the transmembrane region are important for ExbD activity. J Bacteriol. 1996;178:2836-2845
38. Davidson AL, Dassa E, Orelle C, Chen J. Structure, function, and evolution of bacterial ATP-binding cassette systems. Microbiol Mol Biol Rev. 2008;72:317-364
39. Carvajal JJ, Pook MA, dos Santos M, Doudney K, Hillermann R, Minogue S, Williamson R, Hsuan JJ, Chamberlain S. The Friedreich's ataxia gene encodes a novel phosphatidylinositol-4-phosphate 5-kinase. Nat Genet. 1996;14:157-162
40. Prince RC, Grossman MJ. Novel iron-sulfur clusters. TIBS. 1993;18:153-154
41. Demple B, Hidalgo E, Ding H. Transcriptional regulation via redox-sensitive iron-sulphur centres in an oxidative stress response. Biochem Soc Symp. 1999;64:119-128
42. Khoroshilova N, Popescu C, Münck E, Beinert H, Kiley PJ. Iron-sulfur cluster disassembly in the FNR protein of Escherichia coli by O2: [4Fe-4S] to [2Fe-2S] conversion with loss of biological activity. Proc Natl Acad Sci U S A. 1997;94:6087-6092
43. Hobman JL. MerR family transcription activators: similar designs, different specificities. Mol Microbiol. 2007;63:1275-1278
44. Lund PA, Ford SJ, Brown NL. Transcriptional regulation of the mercury-resistance genes of transposon. J Gen Microbiol. 1986;132:465-480
45. O'Halloran TV, Frantz B, Shin MK, Ralston DM, Wright JG. The MerR heavy metal receptor mediates positive activation in a topologically novel transcription complex. Cell. 1989;56:119-129
46. Ma Z, Jacobsen F, Giedroc D. Metal transporters and metal sensors: How coordination chemistry controls bacterial metal homeostasis. Chem Rev. 2009;109:4644-4681
47. Brown NL, Stoyanov JV, Kidd SP, Hobman JL. The MerR family of transcriptional regulators. FEMS Microbiol Rev. 2003;27:145-163
48. Lundemose AG, Kay JE, Pearce JH. Chlamydia trachomatis Mip-like protein has peptidyl-prolyl cis/trans isomerase activity that is inhibited by FK506 and rapamycin and is implicated in initiation of chlamydial infection. Mol Microbiol. 1993;7:777-783
49. Masuzawa T, Sawaki K, Nagaoka H, Akiyama M, Hirai K, Yanagihara Y. Relationship between pathogenicity of Coxiella burnetii isolates and gene sequences of the macrophage infectivity potentiator (Cbmip) and sensor-like protein (qrsA). FEMS Microbiol Lett. 1997;154:201-205
50. Norville IH, Harmer NJ, Harding SV, Fischer G, Keith KE, Brown KA, Sarkar-Tyson M, Titball RW. A Burkholderia pseudomallei macrophage infectivity potentiator-like protein has rapamycin-inhibitable peptidylprolyl isomerase activity and pleiotropic effects on virulence. Infect Immun. 2011;79:4299-4307
51. Pereira PJ, Vega MC, González-Rey E, Fernández-Carazo R, Macedo-Ribeiro S, Gomis-Rüth FX, González A, Coll M. Trypanosoma cruzi macrophage infectivity potentiator has a rotamase core and a highly exposed alpha-helix. EMBO Rep. 2002;3:88-94
52. Cavalieri SJ, Bohach GA, Snyder IS. Escherichia coli alpha-hemolysin: characteristics and probable role in pathogenicity. Microbiol Rev. 1984;48:326-343
53. Forestier C, Welch RA. Identification of RTX toxin target cell specificity domains by use of hybrid genes. Infect Immunol. 1991;59:4212-4220
54. Gadeberg OV, Orskov I, Rhodes JM. Cytotoxic effect of an alpha-hemolytic Escherichia coli strain on human blood monocytes and granulocytes in vitro. Infect Immun. 1983;41:358-364
55. Keane WF, Welch R, Gekker G, Peterson PK. Mechanism of Escherichia coli alpha-hemolysin-induced injury to isolated renal tubular cells. Am J Pathol. 1987;126:350-357
56. Wiles TJ, Dhakal BK, Eto DS, Mulvey MA. Inactivation of host Akt/protein kinase B signaling by bacterial pore-forming toxins. Mol Biol Cell. 2008;19:1427-1438
57. Könninger UW, Hobbie S, Benz R, Braun V. The haemolysin-secreting ShlB protein of the outer membrane of Serratia marcescens: determination of surface-exposed residues and formation of ion-permeable pores by ShlB mutants in artificial lipid bilayer membranes. Mol Microbiol. 1999;32:1212-1225
58. Schönherr R, Hilger M, Broer S, Benz R, Braun V. Interaction of Serratia marcescens hemolysin (ShlA) with artificial and erythrocyte membranes. Eur J Biochem. 1994;15:655-663
59. Wood GE, Dutro SM, Totten PA. Target cell range of Haemophilus ducreyi hemolysin and its involvement in invasion of human epithelial cells. Infect Immun. 1999;67:3740-3749
60. Weiser JN, Chong STH, Greenberg D, Fong W. Identification and characterization of a cell envelope protein of Haemophilus influenzae contributing to phase variation in colony opacity and nasopharyngeal colonization. Mol Microbiol. 1995;17:555-564
61. Blake MS, Gotschlich EC. Purification and partial characterization of the opacity-associated proteins of Neisseria gonorrhoeae. J Exp Med. 1984;159:452-462
62. Arcus VL, Backbro K, Roos A, Daniel EL, Baker EN. Distant structural homology leads to the functional characterization of an archaeal PIN domain as an exonuclease. J Biol Chem. 2004;279:16471-16478
63. Buts L, Lah J, Dao-Thi MH, Wyns L, Loris R. Toxin-antitoxin modules as bacterial metabolic stress managers. TIBS. 2005;30:672-679
64. Zielenkiewicz U, Ceglowski P. Mechanisms of plasmid stable maintenance with special focus on plasmid addiction systems. Acta Biochim Pol. 2001;48:1003-1023
65. Pandey DP, Gerdes K. Toxin-antitoxin loci are highly abundant in free-living but lost from host-associated prokaryotes. Nucleic Acids Res. 2005;33:966-976
66. Mittenhuber G. Occurrence of mazEF-like antitoxin/toxin systems in bacteria. J Mol Microbiol Biotechnol. 1999;1:295-302
67. Gerdes K. Toxin-antitoxin modules may regulate synthesis of macromolecules during nutritional stress. J Bacteriol. 2000;182:561-572
68. Makarova KS, Grishin NV, Koonin EV. The HicAB cassette, a putative novel, RNA-targeting toxin-antitoxin system in archaea and bacteria. Bioinformatics. 2006;22:2581-2584
69. Giguere S, Hondalus MK, Yager JA, Darrah P, Mosser DM, Prescott JF. Role of the 85-kilobase plasmid and plasmid-encoded virulence-associated protein A in intracellular survival and virulence of Rhodococcus equi. Infect Immun. 1999;67:3548-3557
70. Jain S, Bloom BR, Hondalus MK. Deletion of vapA encoding Virulence Associated Protein A attenuates the intracellular actinomycete Rhodococcus equi. Mol Microbiol. 2003;50:115-128
71. Hondalus MK, Mosser DM. Survival and replication of Rhodococcus equi in macrophages. Infect Immun. 1994;62:4167-4175
72. Hondalus MK. Pathogenesis and virulence of Rhodococcus equi. Vet Microbiol. 1997;56:257-268
73. Benoit S, Benachour A, Taouji S, Auffray Y, Hartke A. H2O2 which causes macrophage-related stress, triggers induction of expression of virulence-associated plasmid determinants in Rhodococcus equi. Infect Immun. 2002;70:3768-3776
74. Feldgarden M, Riley MA. The phenotypic and fitness effects of colicin resistance in Escherichia coli K-12. Evolution. 1999;53:1019-1027
75. Cascales E, Buchanan SK, Duché D, Kleanthous C, Lloubès R, Postle K, Riley M, Slatin S, Cavard D. Colicin biology. Microbiol Mol Biol Rev. 2007;71:158-229
76. Johnson WM, Lior H. A new heat-labile cytolethal distending toxin (CLDT) produced by Campylobacter spp. Microb Pathog. 1988;4:115-126
77. Pickett CL, Cottle DL, Pesci EC, Bikah G. Cloning, sequencing, and expression of the Escherichia coli cytolethal distending toxin genes. Infect Immun. 1994;62:1046-1051
78. Cope LD, Lumbley S, Latimer JL, Klesney-Tait J, Stevens MK, Johnson LS, Purven M, Munson RS Jr, Lagergard T, Radolf JD, Hansen EJ. A diffusible cytotoxin of Haemophilus ducreyi. Proc Natl Acad Sci USA. 1997;94:4056-4061
79. Mayer MP, Bueno LC Hansen EJ, DiRienzo JM. Identification of a cytolethal distending toxin gene locus and features of a virulence-associated region in Actinobacillus actinomycetemcomitans. Infect Immun. 1999;67:1227-1237
80. Okuda J, Kurazono H, Takeda Y. Distribution of the cytolethal distending toxin A gene (cdtA) among species of Shigella and Vibrio, and cloning and sequencing of the cdt gene from Shigella dysenteriae. Microb Pathog. 1995;18:167-172
81. Young VB, Knox KA, Schauer DB. Cytolethal distending toxin sequence and activity in the enterohepatic pathogen Helicobacter hepaticus. Infect Immun. 2000;68:184-191
82. Jinadasa RN, Bloom SE, Weiss RS, Duhamel GE. Cytolethal distending toxin: a conserved bacterial genotoxin that blocks cell cycle progression, leading to apoptosis of a broad range of mammalian cell lineages. Microbiology. 2011;157:1851-1875
83. Elwell C, Dreyfus LA. DNase I homologous residues in CdtB are critical for cytolethal distending toxin-mediated cell cycle arrest. Mol Microbiol. 2000;37:952-963
84. Teter K, Allyn R, Jobling M, Holmes R. Transfer of the cholera toxin A1 polypeptide from the endoplasmic reticulum to the cytosol is a rapid process facilitated by the endoplasmic reticulum-associated degradation pathway. Infect Immun. 2002;70:6166-6171
85. Matsuda M, Shigematsu M, Tazumi A, Sekizuka T, Takamiya S, Millar BC, Taneike I, Moore JE. Cloning and structural analysis of the full-length cytolethal distending toxin (cdt) gene operon from Campylobacter lari. Br J Biomed Sci. 2008;65:195-199
86. Dreyfus LA. Cyotlethal Distending Toxin. In Bacterial Protein Toxins. Edited by Burns DL,Barbieri JT, Iglewski BH, Rappuoli. R Washington, DC: ASM Press. 2003:257-270
87. Abuoun M, Manning G, Cawthraw SA, Ridley A, Ahmed IH, Wassenaar TM, Newell DG. Cytolethal distending toxin (CDT)-negative Campylobacter jejuni strains and anti-CDT neutralizing antibodies are induced during human infection but not during colonization in chickens. Infect Immun. 2005;73:3053-3062
88. Xu Z, Yue M, Zhou R, Jin Q, Fan Y, Bei W, Chen H. Genomic characterization of Haemophilus parasuis SH0165, a highly virulent strain of serovar 5 prevalent in China. PLoS One. 2011;6(5):e19631. doi: 10.1371/journal.pone.0019631
Author contact
Corresponding author: Paulraj K Lawrence, Ph.D. Newport Laboratories Inc., 1520 Prairie Drive, Worthington, MN-56187. Email: plawrencecom Phone: 507-372-3020.

 Global reach, higher impact
Global reach, higher impact