ISSN: 1839-9940
J Genomics 2019; 7:56-59. doi:10.7150/jgen.38149 This volume Cite
Research Paper
The Draft Genome of a Hydrogen-producing Cyanobacterium, Arthrospira platensis NIES-46
Center for Biology and Environmental Studies, National Institute for Environmental Studies, 16-2 Onogawa, Tsukuba, Ibaraki, Japan.
Received 2019-7-4; Accepted 2019-8-5; Published 2019-9-18
Abstract
Arthrospira is an economically important cyanobacterium that contains many useful products, including proteins, vitamins, lipids, and pigments, and it is distributed in several alkaline soda lakes. Arthrospira platensis NIES-46 produces large amounts of hydrogen. In this study, we sequenced the NIES-46 draft genome and performed comparative analyses among Arthrospira species to elucidate the genomic background of this strain. The genome consists of 5.7 Mbp with a GC% of 44.5% and encodes 5,008 proteins. Our phylogenetic analysis using multiple orthologous proteins shows that Arthrospira is divided into two clades and that NIES-46 is closely related to A. platensis NIES-39. The genome structure and protein functions are highly conserved between A. platensis NIES-39 and NIES-46, suggesting that these two strains have recently diverged. Genes involved in hydrogen production are well-conserved among Arthrospira species, indicating conserved abilities to produce hydrogen.
Keywords: Arthrospira platensis NIES-46, filamentous cyanobacterium, genome, hydrogen production, phylogeny, Spirulina
Introduction
The filamentous, non-N2-fixing cyanobacterium Arthrospira is an economically important organism that contains a large number of proteins, vitamins, lipids, and pigments, and it is widely used in human and animal food [1]. The natural habitat of Arthrospira is alkaline soda lakes such as Lake Chad in Chad, Lake Texcoco in Mexico, and Lake Chenghai in China; therefore, Arthrospira can tolerate high pH and salt environments [2]. These are useful characteristics for mass-cultivation in open ponds because these conditions can be toxic for contaminating species and predators.
In addition to some nutrients and other useful products, Arthrospira can produce hydrogen via a bidirectional [NiFe]-hydrogenase, an enzyme that catalyses the reversible reduction of protons to H2 coupled to NAD+ regeneration [3]. This enzyme is composed of five subunits encoded by the hox gene cluster (hoxEFUHY). Accessory proteins (encoded by the hypABCDEF gene cluster) are required for the maturation of the active site within the catalytic cluster of the hydrogenase subunit [3]. Several cyanobacterial genomes possess the hox and hyp gene clusters [3]. In particular, Arthrospira maxima is one of the most efficient hydrogen-producing cyanobacterium [4,5], and Arthrospira platensis also produces large amounts of hydrogen [6]. In A. platensis NIES-46, detailed physiology and culture conditions for hydrogen production have been reported [7]. NIES-46 produces hydrogen under dark anaerobic conditions and is enhanced by nitrogen-starvation. However, the genome sequence of NIES-46 is unavailable.
To date, the complete genomes of nine Arthrospira strains have been sequenced, including A. maxima, A. platensis, and unidentified Arthrospira species (https://www.ncbi.nlm.nih.gov/genomes/GenomesGroup.cgi?taxid=35823). These genomes have many repetitive sequences (e.g., group II introns) and a wide variety of restriction-modification systems [8-10]. The repetitive sequences underlie the high genome plasticity observed in this genus [9]. Arthrospira genomes also encode NapA-type Na+/H+ antiporter homologs, which can be involved in salt tolerance in alkaline habitats [11]. The Arthrospira species with available genomes were sampled from lakes or ponds in Africa (NIES-39 and UTEX2342), Europe (Paraca and O9.13F), and Asia (TJSD091 and TJSD092), while the sampling locations of two strains (PCC8005 and PCC9438) are unknown. No genomes of Arthrospira strains sampled on the American continent have been sequenced, and such an undertaking could be interesting for gaining more insight into Arthrospira evolution based on having a more complete picture of its global distribution. In this study, we sequenced the genome of A. platensis NIES-46, which was sampled from Lake Texcoco in Mexico.
Material and Methods
Arthrospira platensis NIES-46 was maintained in 10 mL SOT medium [12] at 20ºC under a 10 h:14 h light:dark cycle. The cells were collected via gentle centrifugation. DNA was extracted using NucleoBond Buffer Set III and NucleoBond AXG 100 (Macherey-Nagel, Düren, Germany), following the manufacturer's protocol. The DNA was sheared into fragments of about 550 bp using a Covaris M220 ultrasonicator (Covaris, Woburn, MA). The DNA library for the MiSeq sequencing was prepared using the NEBNext Ultra II DNA Library Prep Kit for Illumina (New England Biolabs, Ipswich, MA). Paired-end reads (257,288,461 bp) were obtained on the MiSeq platform (Illumina, San Diego, CA) with the 600-cycle MiSeq Reagent Kit v3. The raw reads were trimmed, assembled, and error-corrected using Trimmomatic 0.38 [13], SPAdes v3.11.1 [14], and Pilon v1.22 [15], respectively, all of which were implemented in Shovill v1.0.4 (https://github.com/tseemann/shovill). After removing the short reads (< 200 bp), a gene model construction was generated using the DFAST legacy server [16]. Functional annotation was performed using the EggNOG web server [17]. The draft genome was deposited to the DDBJ/Genbank/ENA repositories with the accession numbers BIMW01000001-BIMW01000343. For phylogenetic analysis, 126 conservative orthologs were used. Lyngbya sp. PCC8106 and Trichodesmium erythraeum IMS101 were used as an out group based on a previous report [8]. The orthologs were determined via reciprocal BLAST best-hit analyses with the following cut-offs: similarity >80% and HSP coverage >80%. Model tests and maximum-likelihood analyses with 200 non-parametric bootstrap replications were performed via IQ-TREE 1.7 [25].
A. platensis NIES-46 genome features
| Feature | Characteristic |
|---|---|
| Assembly length | 5,728,646 bp |
| Number of contigs | 343 |
| N50 | 153 kbp |
| GC content | 44.50% |
| Number of protein-coding genes | 5,008 |
| Number of rRNAs | 4 |
| Number of tRNAs | 38 |
| Number of CRISPRs | 8 |
Results and Discussion
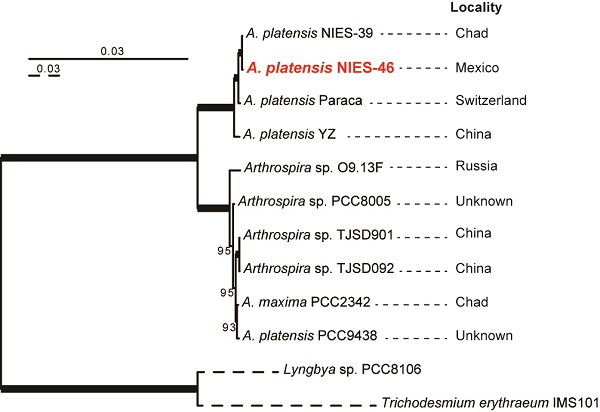
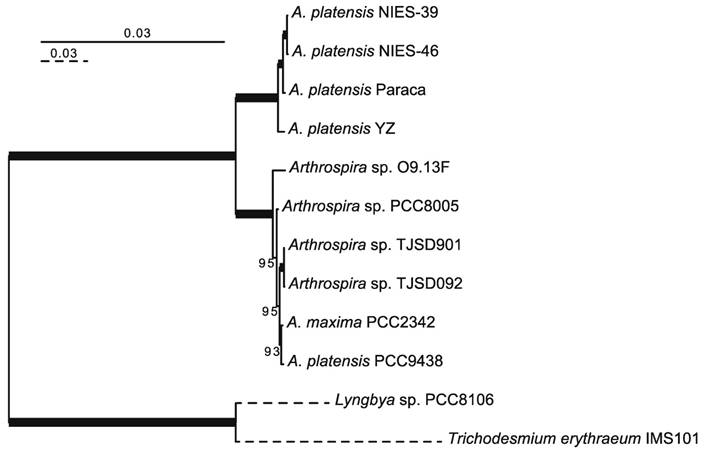
The NIES-46 draft genome is composed of 343 contigs and 5,728,646 bp. The genome completeness is 100% based on a comparison with cyanobacterial marker genes using checkM analysis [19]. The GC% of the genome is 44.5%, and it encodes 5,008 proteins, four rRNAs, 38 tRNAs, and eight CRISPR clusters (Table 1). To infer the phylogenetic relationships between NIES-46 and other Arthrospira strains, we performed a phylogenetic analysis using 126 conservative orthologs (Figure 1). The operation taxonomic units were clearly divided into two clades (bootstrap percentage [BP] = 100): clade A (NIES-39, NIES-46, Paraca, and YZ) and clade B (O9.13F, PCC8005, TJSD091, TJSD092, UTEX-2342, and PCC9438). These clades may correspond to two previously reported clusters determined via rDNA restriction analysis [20] and phylogenetic analysis using the 16S rRNA and cpcBA-IGS sequences [21]. In clade A, NIES-46 was a robust sister of NIES-39 (BP = 100). This observation is surprising because the sampling locations of NIES-46 (Mexico) and NIES-39 (Chad) are separated by the Atlantic Ocean. Although the distribution pattern remains unknown, the strain might be migrated via wind, birds [22,23], or human activities (e.g., through fish stocking) [24]. To elucidate the nucleotide-level similarity between NIES-46 and NIES-39, we performed BLASTn analyses using transcript sequences from both strains with an e-value cut-off <1E-5. The transcript similarity was 99.8% on average. The genome structures of NIES-46 and NIES-39 were compared via progressiveMauve [25]. At least six locally colinear blocks and only two short inversions (9,696 bp and 25,438 bp) were found (Figure S1), contrast to the known high genome plasticity in Arthrospira strains, which includes many rearrangements [8]. These observations suggest that these strains diverged recently. The putative protein functions among the Arthrospira species in clade A were compared based on COG classification [26]. The protein functions encoded by the NIES-46 genome are highly similar to those in the genomes of NIES-39 and the Paraca strain (Figure S2). The similarity of the NIES-46 and NIES-39 genomes is consistent with their shared phenotypic features, including their similar growth conditions [27]. In particular, we focused on genes involved in hydrogen production, including hoxEFUHY and hypABCDEF. The NIES-46 genome encodes all of these genes, whose loci are conserved among Arthrospira strains, suggesting that Arthrospira strains have a conserved hydrogen production ability. Recently, an effective transformation system for A. platensis has been developed using a type 1 restriction inhibitor and liposomes to protect the exogenous DNA [28], and the hox and hyp loci could be future targets for transformation to increase hydrogen productivity.
The NIES-46 genome sequence and its annotation should be useful tools for bioengineering applications, including the generation of mutant strains with increased hydrogen production; furthermore, this study provides genomic insight into the dispersion of Arthrospira species. Because the natural habitats of NIES-46 and NIES-39, Lake Texcoco [29] and Lake Chad [30], respectively, are shrinking, these strains should be preserved in multiple culture collections to sustain Arthrospira genetic diversity.
Abbreviations
BP: bootstrap percentage.
Supplementary Material
Supplementary figures.
Acknowledgements
We thank Nobuyoshi Nakajima (NIES) for the genome sequencing. This work was partially supported by the National BioResource Project for Algae under grant number 17km0210116j0001, which is funded by the Japan Agency for Medical Research and Development (AMED).
Data Deposition
The draft genome was deposited to the DDBJ/Genbank/ENA repositories with the accession numbers BIMW01000001-BIMW01000343.
Competing Interests
The authors have declared that no competing interest exists.
Phylogenetic tree of Arthrospira species. A maximum-likelihood tree was inferred using 126 orthologous proteins. The bootstrap percentages (BPs) are indicated on the nodes. The bold lines represent 100% BP. The dotted branches are shown in quarter-length.
References
1. Habib MAB, Parvin M, Huntington TC, Hasan MR. A Review on culture, production and use of Spirulina as food for humans and feeds for domestic animals and fish. Rome, Italy: Food and Agriculture Organization of the United Nations. 2008
2. Vonshak A. Spirulina: growth, physiology and biochemistry. In: (ed.) Vonshak A. Spirulina platensis (Arthrospira). London, England: CRC Press. 2014:43-65
3. Tamagnini P, Leitão E, Oliveira P. et al. Cyanobacterial hydrogenases: diversity, regulation and applications. FEMS Microbiol Rev. 2007;31:692-720
4. Ananyev G, Carrieri D, Dismukes GC. Optimization of metabolic capacity and flux through environmental cues to maximize hydrogen production by the cyanobacterium “Arthrospira (Spirulina) maxima.” Appl Environ Microbiol. 2008; 74: 6102-6113.
5. Krishnan A, Qian X, Ananyev G, Lun DS, Dismukes GC. Rewiring of cyanobacterial metabolism for hydrogen production: synthetic biology approaches and challenges. In: (ed.) Zhang W, Song X. Synthetic Biology of Cyanobacteria. Advances in Experimental Medicine and Biology. Singapore: Springer. 2018:171-213
6. Ainas M, Hasnaoui S, Bouarab R, Abdi N, Drouiche N, Mameri N. Hydrogen production with the cyanobacterium Spirulina platensis. Int J Hydrogen Energy. 2017;42:4902-4907
7. Aoyama K, Uemura I, Miyake J, Asada Y. Fermentative metabolism to produce hydrogen gas and organic compounds in a cyanobacterium, Spirulina platensis. J Ferment Bioeng. 1997;83:17-20
8. Horikawa H, Fujita N, Okamoto S. et al. Genomic structure of an economically important cyanobacterium, Arthrospira (Spirulina) platensis NIES-39. DNA Res. 2010;17:85-103
9. Xu T, Qin S, Hu Y. et al. Whole genomic DNA sequencing and comparative genomic analysis of Arthrospira platensis: high genome plasticity and genetic diversity. DNA Res. 2016;23:325-338
10. Janssen PJ, Morin N, Mergeay M. et al. Genome sequence of the edible cyanobacterium Arthrospira sp. PCC 8005. J Bacteriol. 2010;192:2465-2466
11. Cheevadhanarak S, Paithoonrangsarid K, Prommeenate P. et al. Draft genome sequence of Arthrospira platensis C1 (PCC9438). Stand Genomic Sci. 2012;6:43-53
12. Ogawa T, Terui G. Studies on the growth of Spirulina platensis. (I) On the pure culture of Spirulina platensis. J Ferment Technol. 1970;48:361-367
13. Bolger AM, Lohse M, Usadel B. Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics. 2014;30:2114-2120
14. Bankevich A, Nurk S, Antipov D. et al. SPAdes: a new genome assembly algorithm and its applications to single-cell sequencing. J Comput Biol. . 2012;19:455-477
15. Walker BJ, Abeel T, Shea T. et al. Pilon: an integrated tool for comprehensive microbial variant detection and genome assembly improvement. PLoS One. 2014;9:e112963
16. Tanizawa Y, Fujisawa T, Nakamura Y. DFAST: a flexible prokaryotic genome annotation pipeline for faster genome publication. Bioinformatics. 2018;34:1037-1039
17. Huerta-Cepas J, Forslund K, Coelho LP. et al. Fast genome-wide functional annotation through orthology assignment by eggNOG-Mapper. Mol Biol Evol. 2017;34:2115-2122
18. Nguyen L-T, Schmidt HA, von Haeseler A, Minh BQ. IQ-TREE: a fast and effective stochastic algorithm for estimating maximum-likelihood phylogenies. Mol Biol Evol. 2015;32:268-274
19. Parks DH, Imelfort M, Skennerton CT, Hugenholtz P, Tyson GW. CheckM: assessing the quality of microbial genomes recovered from isolates, single cells, and metagenomes. Genome Res. 2015;25:1043-1055
20. Belay A, Scheldeman P, Baurain D. et al. Arthrospira ('Spirulina') strains from four continents are resolved into only two clusters, based on amplified ribosomal DNA restriction analysis of the internally transcribed spacer. FEMS Microbiol Lett. 1999;172:213-222
21. Choi G-G, Ahn C-Y, Oh H-M. Phylogenetic relationships of Arthrospira strains inferred from 16S rRNA gene and cpcBA-IGS sequences. Algae. 2012;27:75-82
22. Sukenik A, Hadas O, Kaplan A, Quesada A. Invasion of Nostocales (cyanobacteria) to subtropical and temperate freshwater lakes - physiological, regional, and global driving forces. Front Microbiol. 2012;3:1-9
23. van Leeuwen CHA, van der Velde G, van Groenendael JM, Klaassen M. Gut travellers: internal dispersal of aquatic organisms by waterfowl. J Biogeogr. 2012;39:2031-2040
24. Padilla DK, Williams SL. Beyond ballast water : aquarium and species in aquatic ecosystems. Front Ecol Environ. 2004;2:131-138
25. Darling AE, Mau B, Perna NT. progressiveMauve: multiple genome alignment with gene gain, loss and rearrangement. PloS One. 2010;5:e11147
26. Tatusov RL, Fedorova ND, Jackson JD. et al. The COG database: an updated version includes eukaryotes. BMC bioinformatics. 2003;4:41
27. Kim C-J, Jung Y-H, Oh H-M. Factors indicating culture status during cultivation of Spirulina (Arthrospira) platensis. J Microbiol. 2007;45:122-127
28. Jeamton W, Dulsawat S, Tanticharoen M, Vonshak A, Cheevadhanarak S. Overcoming intrinsic restriction enzyme barriers enhances transformation efficiency in Arthrospira platensis C1. Plant Cell Physiol. 2017;58:822-830
29. Alcocer J, Williams WD. Historical and recent changes in Lake Texcoco, a saline lake in Mexico. Int J Salt Lake Res. 1996;5:45-61
30. Gao H, Bohn TJ, Podest E, McDonald KC, Lettenmaier DP. On the causes of the shrinking of Lake Chad. Environ Res Lett. 2011;6:034021
Author contact
![]() Corresponding author: e-mail address: suzuki.shigekatsugo.jp; postal address: 16-2 Onogawa, Tsukuba, Ibaraki 305-8506 Japan.
Corresponding author: e-mail address: suzuki.shigekatsugo.jp; postal address: 16-2 Onogawa, Tsukuba, Ibaraki 305-8506 Japan.

 Global reach, higher impact
Global reach, higher impact