ISSN: 1839-9940
J Genomics 2017; 5:12-15. doi:10.7150/jgen.17863 This volume Cite
Research Paper
Genomic characterization of eight Ensifer strains isolated from pristine caves and a whole genome phylogeny of Ensifer (Sinorhizobium)
1. School of Science, Monash University Malaysia, Bandar Sunway, Selangor, Malaysia.
2. Genomics Facility, Tropical Medicine and Biology Platform, Monash University Malaysia, Bandar Sunway, Selangor, Malaysia.
3. Department of Biology, University of Akron, Akron, Ohio, USA.
4. Thomas H. Gosnell School of Life Sciences, Rochester Institute of Technology Rochester, NY, USA.
# equal contribution
Published 2017-1-18
Abstract
A total of eight Ensifer sp. strains were isolated from two pristine cave environments. One strain was isolated from a cave water pool located in the Wind Cave National Park, South Dakota, USA and the remaining seven strains were isolated from Lechuguilla Cave of Carlsbad Caverns National Park, New Mexico, USA. Whole genome sequencing and comparative genomic analyses of the eight isolates compared to various type strains from the genera Ensifer and Sinorhizobium demonstrates that although members in these genera can be phylogenetically separated into two distinct clades, the percentage of conserved proteins (POCP) between various type strains from Ensifer and Sinorhizobium are consistently higher than 50%, providing strong genomic evidence to support the classification of the genera Ensifer and Sinorhizobium into a single genus.
Keywords: Ensifer sp. Strains, Sinorhizobium
Introduction
The type species for the genus Ensifer, Ensifer adhaerens ATCC 33212 was initially characterized due to its predation activity against other bacteria in addition to its nitrogen-fixing activity under nutrient-limited growth conditions [1]. The nitrogen-fixing genus Sinorhizobium were later found to be synonymous with the Ensifer. In addition, since the genus Ensifer were classified first, this synonymy was initially resolved by re-classifying members of the Sinorhizobium as Ensifer species. However, given that Sinorhizobium sp. were significant members of the rhizosphere, this created great consternation in the rhizobium scientific community [2]. Adding to this confusion was additional analyses that suggested a clear separation of the Ensifer and Sinorhizobium into two separate genera [3]. The recently published method for genus delineation based on protein conservation [4] and the availability of whole genome of various type strains in this group (including the recently published Ensifer adhaerens ATCC 33212) invite a genome-based investigation of the molecular taxonomy of the genera Ensifer and Sinorhizobium [1]. The goals of this study were aimed to: 1) improve taxon sampling of the genus Ensifer by sequencing and annotating eight additional genomes of Ensifer strains isolated from two pristine cave environments, 2) identify putative gene(s) associated with adaptation to cave environments and 3) utilize various genomic/proteomic information to resolve the question of whether the genera Sinorhizobium and Ensifer should be separated into two distinct genera or be combined into a single genus.
Methods
Strain SD006 was isolated from water in Calcite Lake, which is formed where Wind Cave intersects the Madison Aquifer at a depth of -200 m in Wind Cave National Park, South Dakota, USA and maintained on half-strength tryptic soy agar medium (Merck, Germany). The Lechuguilla Cave (LC) strains (LC11, LC13, LC14, LC54, LC163, LC384, and LC499) were isolated from cave dry wall environment in New Mexico as previously described by Bhullar et al., [5]. DNA extraction was performed using E.Z.N.A Tissue DNA Kit (Omega bio-tek, Norcross, GA). The extracted DNA was subsequently processed using Nextera XT (Illumina, San Diego, CA), quantified using Qubit 2.0 (Invitrogen, Waltham, MA) and sequenced on the MiSeq Illumina sequencing platform located at the Monash University Malaysia Genomics Facility.
Illumina adapter removal, whole genome assembly, in-silico scaffolding and gap closing were performed using Trimmomatic v0.35, SPAdes v3.6.2, SSPACE v3.0 and Gapfiller v1.10, respectively [6-9]. Genomic relatedness among different strains was inferred based on average nucleotide identity (ANI) using JSpecies v1.2.1 [10]. PhyloPhlan v0.99 was subsequently used to infer the evolutionary relationship among the sequenced strains and related strains/species [11].
Pangenome analysis was performed with Roary (https://sanger-pathogens.github.io/Roary/) using a protein identity cut-off of 90% for clustering orthologs. Genus delineation was determined based on the percentage of conserved proteins (POCP) as described by Qin et al [4] whereby strains sharing pairwise POCP value of >50% belong to the same prokaryotic genus.
Data description
The genome size of the strains sequenced in this study ranges between 6.0 to 7.5 megabases with N50 and GC content ranging from 180,000 to 232,000 bp and 61.5% to 62.3%, respectively (Table 1). By rooting the constructed tree with several members of the order Rhizobiales as the outgroup, maximum likelihood inference based on the alignment of 400 conserved proteins shows a clear separation of the Ensifer/Sinorhizobium group into two clades (Clade I and Clade II) with maximal local support values inferred by the Shimodaira-Hasegawa test (SH-like local supports). Clade 1 consists of mainly strains from the genus Ensifer including the type strain E. adhaerens. On the other hand, Clade 2 consists of mostly strains from the genus Sinorhizobium including 3 type strains e.g. Sinorhizobium arboris LMG14919, Sinorhizobium saheli LMG7837 and Sinorhizobium fredii USDA205. Ensifer sojae CCBAU05684 is the only strain with Ensifer species designation that demonstrated monophyletic clustering within the “Sinorhizobium” clade. Based on phylogenetic clustering (within Clade 1; Fig. 1C), the cave isolates reported in this study were designated as members of the genus Ensifer. Additionally, these strains represent at least two genospecies of Ensifer distinct from the type species E. adhaerens ATCC 33499, as evidence by their pairwise average nucleotide identity of less than 85% (ANI of >95% indicates identical genospecies; Fig. 1A)[12]. The first cave genospecies consists of strains LC163, LC54, LC384 and SD006 and the second consists of strains LC11, LC14 and LC499.
Genome annotation information for the isolated strains. The table shows the bioproject, genome accession numbers, genome size, GC range (%), N50 range (bp).
| Strain | BioProject | GenBank Accession Number | Genome size (bp) | GC (%) | N50 (bp) |
|---|---|---|---|---|---|
| SD006 | PRJNA288791 | LGAP00000000 | 7,461,094 | 61.53 | 187,147 |
| LC11 | PRJNA328912 | MBSO00000000 | 6,217,316 | 62.21 | 181,097 |
| LC13 | PRJNA328912 | MBSP00000000 | 6,217,834 | 62.21 | 180,792 |
| LC14 | PRJNA328912 | MBSQ00000000 | 6,225,177 | 62.21 | 180,778 |
| LC54 | PRJNA328913 | MBSR00000000 | 6,521,514 | 62.00 | 231,502 |
| LC163 | PRJNA328914 | MBSS00000000 | 7,033,756 | 61.82 | 228,827 |
| LC384 | PRJNA328915 | MBST00000000 | 6,217,481 | 62.22 | 167,160 |
| LC499 | PRJNA328916 | MBSU00000000 | 7,026,316 | 61.82 | 195,448 |
Phylogenetic analysis of cave strains (SD006, LC11, LC13, LC14, LC54, LC163, LC384 and LC499) with other members of Rhizobiales and their genomic similarity. (A) Heatmap showing pair-wise average nucleotide identity based on MUMMER calculation (ANIm) among the cave isolates and Ensifer adhaerans ATCC 33499T. (B) Percentage of conserved proteins (POCP) comparison of strains among the Ensifer/Sinorhizobium clades and strains from the Sphingomonadaceae family designated as “S” in the X-axis. Red horizontal line indicates 50% cutoff value. (C) Maximum likelihood tree of the order Rhizobiales. The tree was rooted using members of the family Sphingomonadaceae as the outgroup. Type strains are indicated by the superscript letter “T”. Values in nodes depict local SH-support and branch length indicates the number of substitution per site.
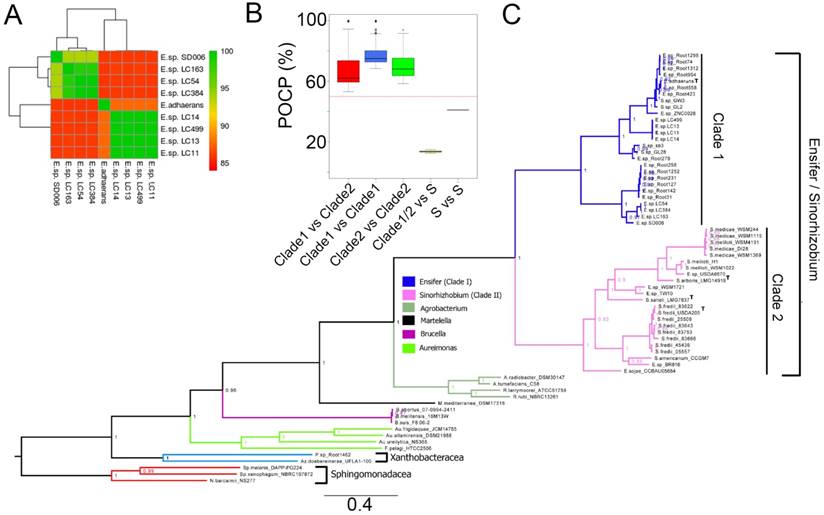
Although the separation of Ensifer and Sinorhizobium into two distinct clades corroborates with previous study by Martens et al. [3] who observed similar separation based on the phylogenetic analysis of ten concatenated house-keeping genes (atpD, dnK, gap, glnA, gltA, gyrB, pnp, recA, rpoB and thrC), all pairwise POCP values among members of Ensifer and Sinorhizobium are consistently higher than 50% (Fig. 1B) thus providing convincing genomic evidence that they represent two major clades within the same genus.
Strain SD006, the only isolate from an aquatic cave environment, has the largest genome size among the eight cave isolates with more than 427,000 bps of additional genomic information than the strain that exhibits the smallest genome (LC11; Table 1). Pan-genome analysis of SD006 and members of its genospecies identified up to 2,106 unique genes. Functional annotation of this unique proteome in SD006 led to the identification of a gene coding for aquaporin (Uniprot entry: A0A0L8BEZ5; locus tag: AC244_32060). The aquaporin protein has been shown to be involved in regulating responses related to changes in environmental osmolality [13] that may be more prevalent in the isolation source of strain SD006 e.g. an aqueous environment, compared to a dry limestone surface where the seven other strains were isolated. The putative aquaporin protein has the highest similarity score of 82.8% (as of 15th August 2016) to Uniprot entry A0A072CG14 from the soil isolate Sinorhizobium americanum CCGM7.
Strain LC11 has been previously demonstrated to exhibit in-vitro predation activity against Micrococcus sp. strain LC524 using methods such as the cross streak and predation activity assay, followed by the visualization of the predation activity using a scanning electron microscopy (SEM) [14]. However, its predation requirement(s) differs substantially from the type strain E. adhaerans ATCC 33499 e.g. while strain LC11 readily tracks prey at pH 8.0, similar to the pH of the cave environment. E. adhaerens ATCC 33499 usually exhibits predation at a more acidic pH of 6.0-6.5.
The availability of whole genome sequences of seven Ensifer sp. will be useful for the identification of genes associated with bacterial predation and/or cave adaptation in members of the genus Ensifer.
Nucleotide sequence accession numbers
The genome sequences of strains described in this study have been deposited at GenBank as described in Table 1. The version described in this paper is the first version.
Acknowledgements
HKSK, WWHE, MHT and HMG thank the Monash University Malaysia Tropical and Medicine Biology for financial and infrastructure support. MAS and AOH acknowledge the College of Science (COS) at the Rochester Institute of Technology (RIT) and The Gosnell School of Life Sciences (GSOLS) at RIT for ongoing support.
Competing Interests
The authors have declared that no competing interest exists.
References
1. Rogel MA, Hernandez-Lucas I, Kuykendall LD, Balkwill DL, Martinez-Romero E. Nitrogen-fixing nodules with Ensifer adhaerens harboring Rhizobium tropici symbiotic plasmids. Applied Environmental Microbiology. 2001;67:3264-8
2. Young JM. Sinorhizobium versus Ensifer: may a taxonomy subcommittee of the ICSP contradict the Judicial Commission? International Journal of Systematic and Evolutionary Microbiology. 2010;60:1711-3
3. Martens M, Dawyndt P, Coopman R, Gillis M, De Vos P, Willems A. Advantages of multilocus sequence analysis for taxonomic studies: a case study using 10 housekeeping genes in the genus Ensifer (including former Sinorhizobium). International Journal of Systematic and Evolulitonary Microbiology. 2008;58:200-14
4. Qin QL, Xie BB, Zhang XY, Chen XL, Zhou BC, Zhou J. et al. A proposed genus boundary for the prokaryotes based on genomic insights. Journal of Bacteriology. 2014;196:2210-5
5. Bhullar K, Waglechner N, Pawlowski A, Koteva K, Banks ED, Johnston MD. et al. Antibiotic resistance is prevalent in an isolated cave microbiome. PLoS One. 2012;7:e34953
6. Boetzer M, Henkel CV, Jansen HJ, Butler D, Pirovano W. Scaffolding pre-assembled contigs using SSPACE. Bioinformatics. 2011;27:578-9
7. Boetzer M, Pirovano W. Toward almost closed genomes with GapFiller. Genome Biology. 2012;13:R56
8. Bolger AM, Lohse M, Usadel B. Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics. 2014;30:2114-20
9. Nurk S, Bankevich A, Antipov D, Gurevich AA, Korobeynikov A, Lapidus A. et al. Assembling single-cell genomes and mini-metagenomes from chimeric MDA products. Journal of Computational Biology. 2013;20:714-37
10. Burall LS, Grim CJ, Mammel MK, Datta AR. Whole Genome Sequence Analysis Using JSpecies Tool Establishes Clonal Relationships between Listeria monocytogenes Strains from Epidemiologically Unrelated Listeriosis Outbreaks. PLoS One. 2016;11:e0150797
11. Segata N, Bornigen D, Morgan XC, Huttenhower C. PhyloPhlAn is a new method for improved phylogenetic and taxonomic placement of microbes. Nature Communication. 2013;4:2304
12. Richter M, Rosselló-Móra R. Shifting the genomic gold standard for the prokaryotic species definition. Proceedings of the National Academy of Sciences of the United States of America. 2009;106:19126-31
13. Takata K, Matsuzaki T, Tajika Y. Aquaporins: water channel proteins of the cell membrane. Progress in histochemistry and cytochemistry. 2004;39:1-83
14. Wilks M. Predation Mediated Carbon Turnover in Nutrient-Limited Cave Environments. University of Akron. 2013
Author contact
Corresponding author: Michael A. Savka, Thomas H. Gosnell School of Life Sciences, College of Science, Rochester Institute of Technology, Rochester, NY, 14623 USA. Email: massbiedu; Office: 585-475-5141

 Global reach, higher impact
Global reach, higher impact