ISSN: 1839-9940
J Genomics 2017; 5:77-82. doi:10.7150/jgen.20216 This volume Cite
Research Paper
A glimpse into the genetic basis of symbiosis between Hydrogenophaga and their helper strains in the biodegradation of 4-aminobenzenesulfonate
1. Genomics Facility, Tropical Medicine and Biology Platform, Monash University Malaysia, Jalan Lagoon Selatan, Bandar Sunway, 47500 Selangor, Malaysia
2. School of Science, Monash University Malaysia, Jalan Lagoon Selatan, Bandar Sunway, 47500 Selangor, Malaysia
3. Centre for Integrative Ecology, School of Life and Environmental Sciences, Deakin University, Pigdons Road, Waurn Ponds, Victoria 3216 Australia
Abstract
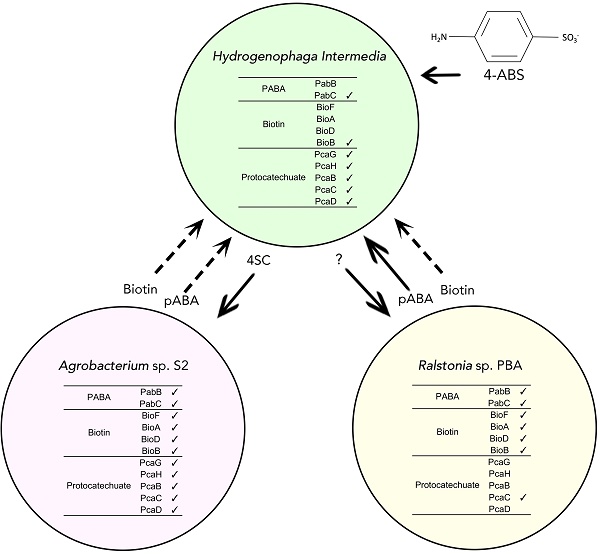
We report the whole genome sequences of Hydrogenophaga intermedia S1 and Agrobacterium radiobacter S2, the first reported bacterial co-culture capable of degrading 4-aminobenzenesulfonate (4-ABS), a recalcitrant industrial waste product. To gain insights into the genetic basis for the syntrophic interaction between this symbiotic pair and also another recently reported Hydrogenophaga associated co-culture, Hydrogenophaga sp. PBC and Ralstonia sp. PBA, we performed detailed genetic analysis of these four strains focusing on the metabolic pathways associated with biotin, para-aminobenzoic acid (pABA), and protocatechuate metabolism. Both assembled Hydrogenophaga draft genomes are missing a majority of the genetic components associated in the biosynthetic pathway of pABA and biotin. Interestingly, a fused pABA synthase was found in R. sp PBA but not in A. radiobacter S2. Furthermore, using whole genome data, the taxonomic classification of R. sp. PBA and A. radiobacter S2 (both previously inferred from 16S rRNA gene) was re-investigated, providing new evidence to propose for their re-classification at the genus and species level, respectively
Keywords: p-aminobenzoic acid, biotin, co-culture, phylogenomics, Hydrogenophaga, Agrobacterium, 4-aminobenzenesulfonate

 Global reach, higher impact
Global reach, higher impact